Proteins Perform Many Functions
The preceding pages describe how the genetic code in DNA specifies the assembly of proteins, which are polypeptides, meaning that their primary structure consists of a long linear assembly of amino acids. As the amino acids are being linked together into a growing polypeptide begins to fold as a result of the interactions among the amino acids. Different segments of a protein can assume different shapes: random coils, helices, and zig-zagging segments that form sheet-like structures; these secondary structures are then folded in ways that establish the tertiary structure, and these fold yet again to create the final quaternary structure, as shown in the graphic below.
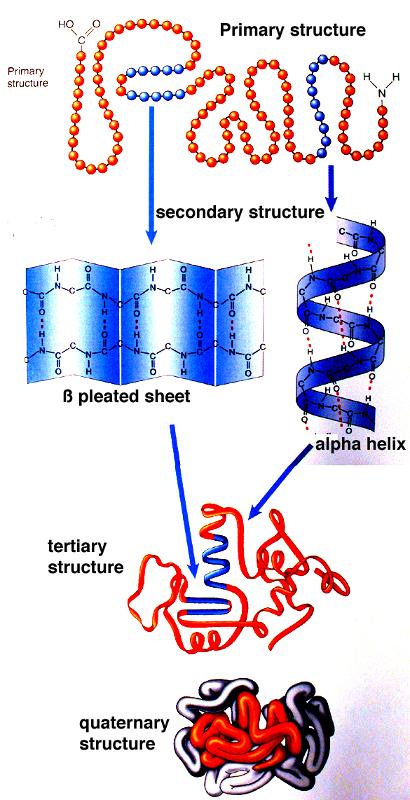
Source: http://academic.brooklyn.cuny.edu/biology/bio4fv/page/3d_prot.htm
Each protein has a specific shape that determines its function. The illustration below summarizes just some of the many functions carried out by proteins.
There are many thousands of enzymes that participate in synthesis and metabolism.
- Antibodies are proteins that are specialized to bind to specific foreign proteins found on viruses and bacteria for example.
- Proteins can act as receptor molecules to which signal molecules can bind to trigger intracellular events
- Proteins act as signal molecules (e.g., insulin for glucose transport or gastrin which stimulates acid secretion in the stomach)
- There are structural proteins (e.g., collagen and elastin)
- Proteins provide the internal structure of all of our cells
- Contractile proteins in skeletal muscles and in the smooth muscle cells in our gastrointestinal tract, respiratory tract, uro-genital tract, and our blood vessels
- Some protein subunits self-associate into more elaborate structures that form conduits across cell membranes for transport of molecules in and out of a cell.
- Proteins serve as shuttles to transport molecules and to transport oxygen
- Proteins can be pigments, such as melanin in our skin
Genetic Variation: Mutations and SNPs
All humans have the same set of genes, and the sequence of our base pairs is remarkably similar. However, this doesn't mean that we all have exactly the same nucleotide sequence in our genome. If this were the case, then all humans would be clones having exactly the same genetic code. The passage of the genetic code from generation to generation via the sperm and ova of our ancestors requires replication of DNA, and while replication is remarkably precise, errors occasionally occur and produce changes in the base sequence. It is these changes that produce the variation that makes each of us genetically unique (except for identical twins), and they also drive evolution of species
Mutations
Mutations are random changes in the sequence of base pairs in DNA, and mutagens are factors that cause mutations, e.g., chemicals or radiation (UV light, x-rays, gamma radiation). Mutagens result four patterns of alteration in the base sequence:
- Replacement (substitution) of a single base pair
- Addition of one or more base pairs
- Deletion of one or more base pairs
- Relocation of a segment of base pairs
The effects of these alterations depend on several factors.
Substitution of One Base Pair
Substitution of one base pair for another produces the smallest change in the code, but it's effects can range from none to massive depending on the details.
The example below shows an initial sequence of base pairs on the left, the mRNA transcript, and the sequence of amino acids that would result. The right portion shows that substitution of thymine (T) for adenine (A) changes the third base in the mRNA transcript, but the amino acid sequence is unchanged, because GUU and GUA are both codons for the amino acid valine. In fact, there are four codons that specify insertion of the amino acid valine: GUU, GUC, GUA, and GUG. If a normal gene had a GUU codon (specifying insertion of valine) at a particular place, changing the third base from uracil (U) to adenine (A) would still produce a protein that had the amino acid valine in the correct position. Consequently, the protein would be unchanged, and the mutation would be "silent."
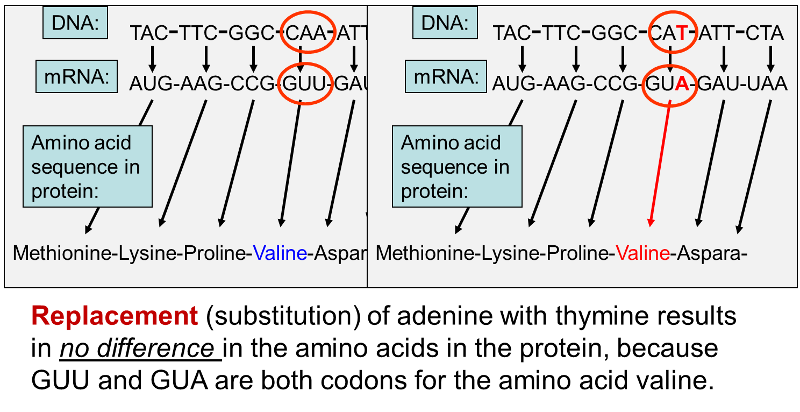
In other cases, however, substitution of a single base pair leads to a change in one of the amino acids in the protein product as shown below.
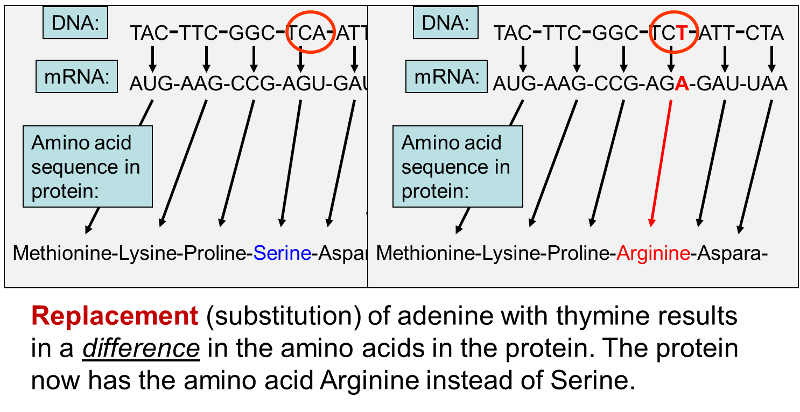
Once again, an adenine (A) in DNA has been replaced by a thymine (T), but in this case the codon in the mRNA changes from AGU, which is a codon for the amino acid serine to AGA , which codes for the amino acid arginine. Changing one amino acid in a protein may have no discernible effect on the protein's function, but at times, changing even a single amino acid can have a massive effect. An example of this is seen with sickle cell anemia.
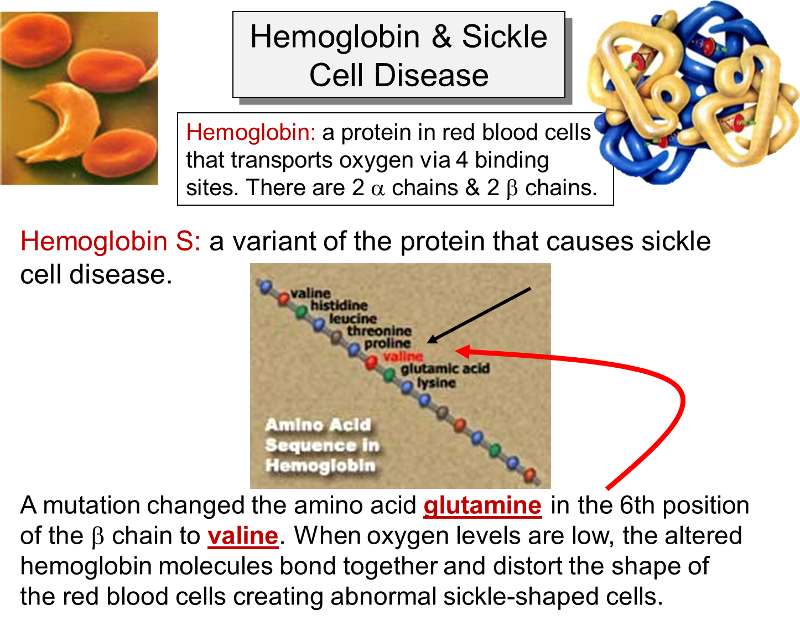
Sickle cell anemia is due to an inherited mutation in the gene that encodes for the β- chain of hemoglobin. Hemoglobin is a protein found in red blood cells which plays a key role in transporting oxygen from the lungs. It is composed of four protein subunits: two α- and two β-chains. In sickle cell anemia the key defect is caused by a mutation that replaces the hydrophilic amino acid glutamic acid (glutamate) with the hydrophobic amino acid valine at the sixth position of the β-chains. This causes the hemoglobin molecules to stick together when oxygen levels are low. As a result, the hemoglobin molecules form long fiber-like chains that distort the shaped of the red blood cells. Red blood cells are normally biconcave disks that flow easily through arteries and capillaries, but sickling causes red blood cells to stick to one another and occlude small arteries causing ischemia in various tissues and organs.
in various tissues and organs.
Link to more information on sickle cell disease.
Addition or Deletion of One Base Pair
In contrast to substitution of a single base, addition or deletion of a single base cause a substantial disruption of the sequence of amino acids in the protein product. This is because the mRNA transcript is read as three-letter codons, and insertion or deletion of a single base causes a frame shift in the sequence that throws off all of the downstream codons. Consider the following hypothetical sequence of DNA and the mRNA transcript it would produce and the final amino acid sequence. For illustration, I begin with triplets with the same base, e.g., AAA, TTT, etc.
DNA -- AAA-CCC-TTT-GGG-GGG- TTT-CCC-TTT-AAA-AAA - etc.
mRNA -- UUU-GGG-AAA-CCC-CCC-AAA-GGG-AAA-UUU-UUU- etc.
Amino Acids -- Phe -- Gly-- Lys--Pro--Pro --Lys --Gly --Lys--Phe--Phe--etc.
Insertion of a single base might result in the following:
DNA -- AAA-ACC-CTT-TGG-GGG-GTT-TCC-CTT-TAA-AAA-A etc.
mRNA --UUU-UGG-GAA-ACC-CCC-CAA-AGG-GAA-AUU-UUU-U- etc.
Amino Acids -- Phe--Trp- Glu--Thr--Pro -Gln -Arg-Glu--Ile--Phe--etc.
The insertion of a base advances all of the letters by one position, but the codons are still read as triplets, so the code is thrown off from the point of the insertion, and most of the amino acids are changed, although occasionally the original amino acid is retained by chance such as the Pro (proline) and the terminal Phe (phenylalanine) in the example above.
Deletion of a single base has the same effect, i.e., it causes a frame shift that changes all of the codons downstream from the point of the deletion.
The short video below illustrates the effects of substitution, insertion, and deletion of single base pairs.
Relocation of a Segment of Nucleotides
The other mechanism that has been described for mutation is the relocation of an entire segment of nucleotides. This is illustrated below starting with the same hypothetical sequence of bases in DNA.
DNA -- AAA-CCC-TTT-GGG-GGG- TTT-CCC-TTT-AAA-AAA - etc.
mRNA -- UUU-GGG-AAA-CCC-CCC-AAA-GGG-AAA-UUU-UUU- etc.
Amino Acids -- Phe -- Gly-- Lys--Pro--Pro --Lys --Gly --Lys--Phe--Phe--etc.
Relocation of a segment of bases might result in:
DNA -- AAA-CCT- CCC-TTT-AAC-TTT-GGG-GGG-TTA-AAA - etc.
mRNA -- UUU-GGA-GGG-AAA-UUG-AAA-CCC - CCC - AAU-UUU- etc.
Amino Acids -- Phe -- Gly-- Lys--Pro--Leu --Phe --Pro -- Pro--Asn--Phe--etc.
This kind of relocation can have variable effects depending on the size of the relocated segment and whether or not it also cause frame shift errors.
Single Nucleotide Polymorphisms (SNPs)
We noted earlier that an error in replication of DNA can result in substitution of one base for another. For example, the human genome might have the following segment of base pairs in double-stranded DNA:
...CGATATTCCTATCGAATGTC....
...GCTATAAGGATAGCTTACAG...
However, a substitution of a single base pair (changing the bold-faced TA pair to GC) might result in a sequence that read as follows:
...CGATAGTCCTATCGAATGTC....
...GCTATCAGGATAGCTTACAG...
We saw earlier that if a substitution like this occurred within a gene, i.e., within a coding area, it could cause a disease like sickle cell anemia, but it is also possible that it might have no discernible effect. Remember, however that coding areas (genes) occupy only 3-5% of our DNA, and substitutions like occur more commonly in the much more extensive non-coding areas between the genes. The accumulation of these random substitutions over time has resulted in many small differences in the sequence of base pairs from person to person. These small differences are found only about once every 300 base pairs on average. However, since we have 3 billion base pairs, this means that there are about 10 million of these small differences in our genome. These small variations have been dubbed "single nucleotide polymorphisms" or SNPs (pronounced "snips"). Each SNP is a difference in a single base in DNA.
In the discussion of mutations we saw that substitution of a single base (a SNP) within a coding area may or may not have an effect on phenotype. In contrast, SNPs occurring in non-coding areas of the genome have no effect on phenotype (characteristics). Nevertheless, each person has a unique SNP pattern, however, and this is potentially useful in a number of ways.
- In order to look for genes that influence the likelihood of a disease, one can take blood samples from a group of individuals with the disease disease and look for similarities in the their SNP patterns. If diseased individuals are found to have a particular SNP, one can then look for differences in SNP patterns in diseased and non-diseased individuals to see if there is an association. Such a study is really just a particular type of case-control study. As a result, it is sometimes possible to establish certain SNP profiles that are associated with a greater likelihood of certain diseases.
- SNPs that may help predict an individual's response to certain drugs, their susceptibility to environmental factors such as toxins.
- SNPs can be used for forensic analysis to match DNA samples.
- SNPs can also determine how closely two individuals are related.
For more information about SNPs go to the two following links:
- http://ghr.nlm.nih.gov/handbook/genomicresearch/snp
- http://www.ncbi.nlm.nih.gov/About/primer/snps.html