
DNA, Genetics, and Evolution
After successfully completing this section, the student will be able to:
> Chromosome
> Gene
> Gene product
> Allele
> Genotype
> Phenotype
> Mitosis
> Meiosis

All living organisms have one or more chromosomes that contain the code that directs the synthesis of proteins that are essential for its structure and function. In bacteria proteins can be structural and they can be enzymes that perform metabolic functions that can breakdown nutrients that provide energy and provide structural building blocks for growth and replication.
Each chromosome is, if fact, an enormous DNA molecule. Molecules are generally so small that they can't be seen even with a microscope, but chromosomes can be seen with a microscope under certain circumstance, particularly when a cell is about to divide. The illustration below shows the 46 chromosomes that contain the human genome.
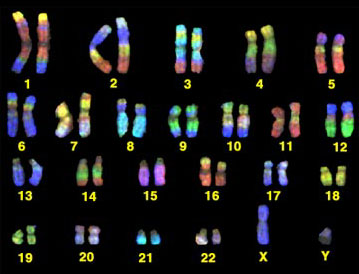
Source: http://www.yorku.ca/kdenning/++2140%202006-7/2140-17oct2006.htm
There are 22 homologous pairs and two sex chromosomes (the X and Y chromosomes). One chromosome in each pair is inherited from one's mother and one from one's father. Each chromosome is a single molecule of DNA. The illustration below illustrates this by imagining that we have grabbed one end of a chromosome and pulled it out to reveal that it is an extremely long polymer consisting of a double helix. In fact, if we were to take a single human chromosome and stretch it out, it would be about 5 centimeters long (about 2 inches), and all 46 chromosomes would be about 2 meters long if they were stretched out and laid end to end. Our cells have all 46 chromosomes, but they are coiled around proteins and highly coiled into the form of the chromosomes that are seen to the right. The chromosomes of eukaryotes are contained within the membrane-bound nucleus.
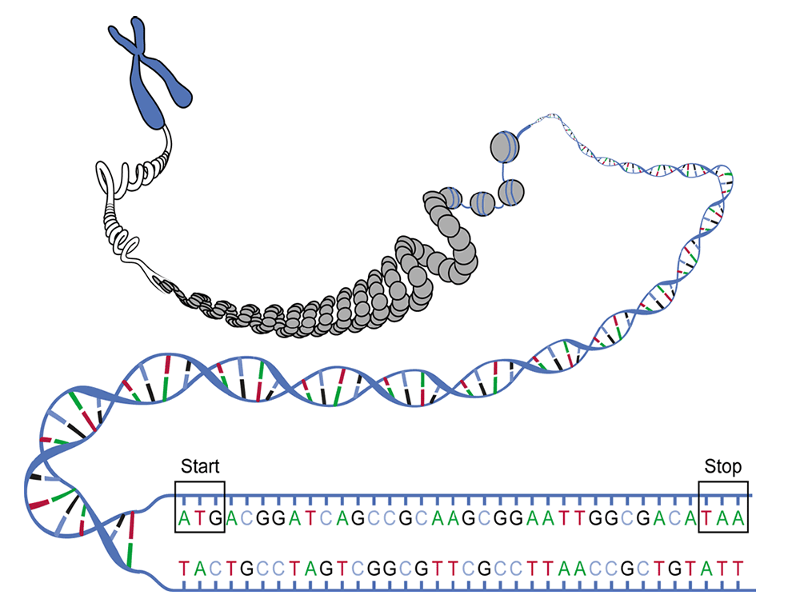
Source: https://biologywarakwarak.wordpress.com/2012/01/15/the-3-magical-rules-to-determine-the-amino-acid-chain-from-a-dna-piece-without-error/
But DNA provides the essential genetic code for all living organisms, including bacteria. The bacterium E. coli has a single circular chromosome (DNA molecule) which is also coiled, supercoiled, and packaged with proteins, but in prokaryotes the chromosome is located in the cytoplasm instead of being contained in a membrane- bound nucleus.
DNA is an abbreviation for deoxyribonucleic acid, which is an extremely long polymer made from units called nucleotides. The illustration below shows the structure of both DNA and RNA (ribonucleic acid.)
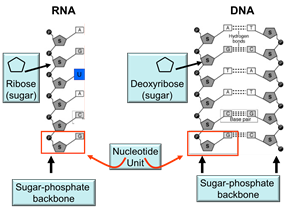
The backbone of each molecule is composed of alternating sugars (the pentagon with the "S") and phosphate groups (shown with "P), and each sugar is also covalently bonded to one of the following nucleotide bases:
A nucleotide "unit" (outlined by the red box in the illustration] consists of a sugar molecule, a phosphate, and one of the five. Consequently, one can think of DNA as an extremely long double-stranded polymer of nucleotides. Note also that the two strands of DNA are held together by hydrogen bonds between complementary bases on the two strands. The figure below demonstrates this complementarity. In DNA the base thymine always bonds to adenine, while cytosine always bonds to guanine because of their complementary chemical structure and "fit". As a result of this complementary structure, if the base sequence of one strand is known, then the structure of the other strand can be deduced.
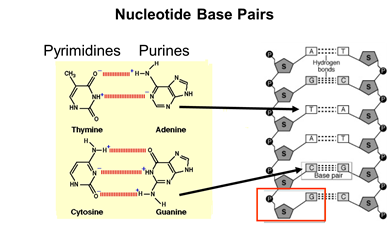
Each of our cells has a complete set of our 46 chromosomes, i.e., our entire genome. Altogether our 46 chromosomes contain about 6 billion nucleotides, i.e., 3 billion base pairs. Each chromosome contains thousands of "genes." The segments of DNA that contain genes (referred to as "coding areas") take up only 3-5% of our DNA; the rest of the DNA consists of " non-coding areas ." Altogether our 23 pairs of chromosomes with their 3 billion base pairs carry the code for 20,000-25,000 genes. Most of the genes are transcribed into "messenger RNAs" (mRNA) that provide a template that is used to translate the code into specific proteins. However, about 100 genes are transcribed into "ribosomal RNAs" and "transfer RNAs" that also play a vital role in the synthesis of proteins, which will be described shortly.
The sequence of bases in DNA can be thought of as the "letters" that provide the basis for the genetic code for all of the proteins synthesized by our bodies, and these, in turn, provide the basis for the structure of all of our cells, all of our enzymes, and all of our inherited traits and characteristics. As noted above, the genetic code is contained in chromosomes which are gigantic molecules of DNA complexed with proteins and wound into a compact structure. Humans have 23 pairs of chromosomes, which carry our entire genome. In eukaryotes chromosomes are located in the cell nucleus, but prokaryotes (bacteria) have a more primitive cellular structure, and they do not have a true nucleus. Instead, the single bacterial chromosome is in the cytoplasm in an area sometimes referred to as the "nucleoid." The production of cellular proteins requires two major processes.
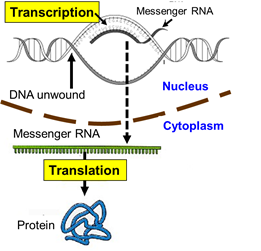
First, cellular signals reaching the nucleus cause the TATA-binding protein to the starting point of a particular gene. Additional transcription factors then bind, and an enzyme called RNA polymerase II then binds to the complex. The the polymerase causes the strands of DNA to separate temporarily, and the enzyme synthesizes a strand of messenger RNA (mRNA) to using the the sequence of bases on one strand of DNA (the coding strand) to create a complementary strand of mRNA.
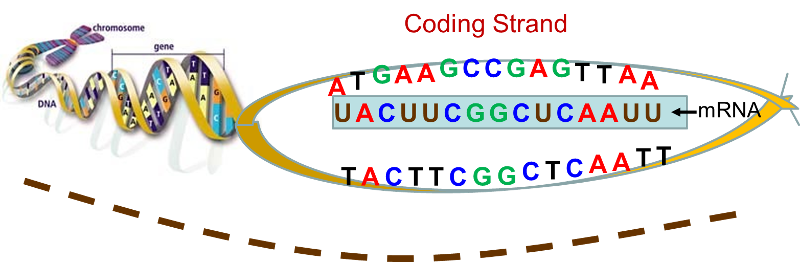
By complementary we mean that the base sequence on mRNA has bases that are the complementary pairs of those on DNA. guanine (G) dictates the insertion of its complement, cytosine (C), and cytosine dictates the insertion of guanine (G); thymine (T) dictates insertion of adenine (A) on mRNA and adenine dictates the insertion of uracil (U). [Note that RNA uses uracil in place of thymine.] Once the strand of mRNA has been created, it leaves the nucleus through pores in the nuclear membrane. The video below gives a fairly detailed picture of the process of transcription. .
Once the mRNA is in the cytoplasm, it binds to a ribosome, which is composed of protein and a different type of RNA called ribosomal RNA (rRNA). One can think of the ribosome as the work bench where protein is synthesized by covalently bonding amino acids in the sequence specified by the code on the mRNA.
One can think of the sequence of bases on mRNA as a series of code letters that are read as a series of three letter "words."
For example, if mRNA had a sequence of bases such as
"...AUGAAGCCGAGUUAAGAU...."
This sequence would, in effect, be read as a series of three-letter words referred to as "codons", each of which specified the insertion of a specific amino acid. In the example just above, the codons or "words" would be:
"...AUG-AAG-CCG-AGU-UAA-GAU...."
Each of these three letter words specifies the insertion of one of the 20 amino acids that make up human proteins. The amino acids are shuttled to the ribosome by a family of transfer RNAs (tRNA), and there are specific tRNAs for each amino acid. The tRNAs consist of a single strand of RNA, but the strand tends to fold back on itself and create loops that are held in place by hydrogen bonds between segments of the tRNA as shown in the illustration below.
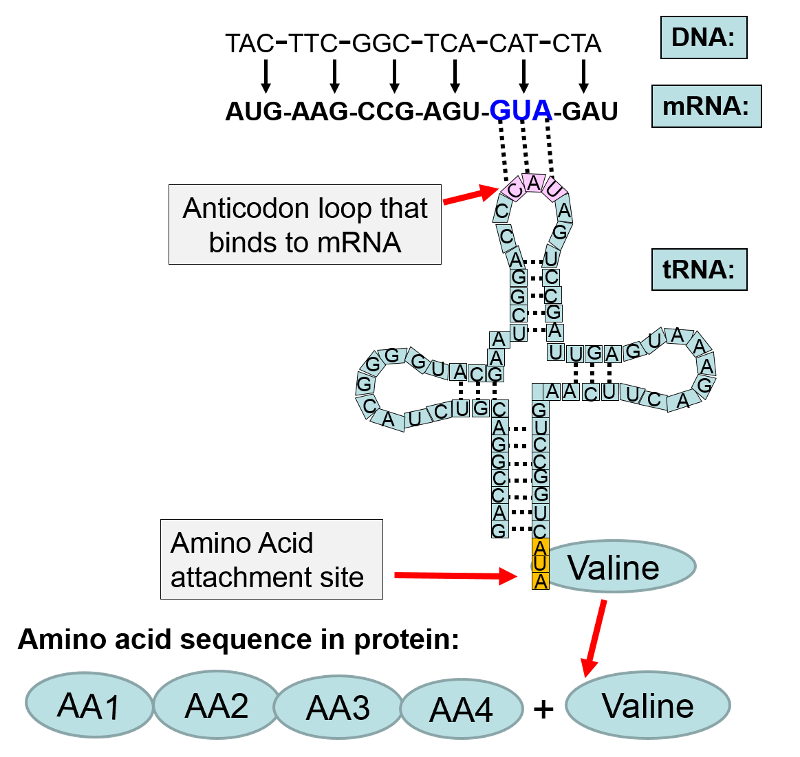
In the illustration above the base sequence CAT on DNA was transcribed to become the codon GUA on messenger RNA. The mRNA left the nucleus and attached to a ribosome where protein synthesis (translation) was initiated. Each codon on mRNA specified a particular amino acid to be added to the growing protein chain. In this example, the first four amino acids are designated as "AA1-AA2-AA3-AA4". The next codon on mRNA was "GUA." The complement to GUA is "CAU" which is the anticodon on a transfer RNA that carries the amino acid valine. The anticodon CAU on the tRNA for valine bonded to the GUA codon on mRNA. This positioned valine as the next amino acid in sequence, and with the addition of cellular energy (ATP), valine became covalently bonded to AA4 in the amino acid chain.
In the section above on transcription, we focused on creating the mRNA for a specific gene; those events took place in the cell nucleus. The figure below illustrates the subsequent events that take place after mRNA leaves the nucleus and attaches to a ribosome and initiates translation.

Sixty-one codons specify an amino acid, and the remaining three act as stop signals for protein synthesis. For example, in the figure below the codon UGA signals an end to synthesis of the protein. The code for all possible three-letter codons on mRNA is shown in the blue table below. Note that there is some redundancy in the code. For example, there are four separate codons for the amino acid proline. Nevertheless, the code is unambiguous, because no triplet codes for more than one amino acid. In addition, with only a few minor exceptions, the same code is universally found in viruses, bacteria, protists, plants, fungi, and animals.
Note that in the example below, UGA, is a signal to STOP, meaning that the amino acid chain is complete and no more amino acids are to be added. Bear in mind that these illustrations include just short sequences of codons, and an actual protein would generally have a much longer sequence. Nevertheless, these examples illustrated how the code is transcribed from DNA to mRNA and how the mRNA is then translated in order to specify the sequence of amino acids in a particular protein which is the product of that particular gene on a chromosome.
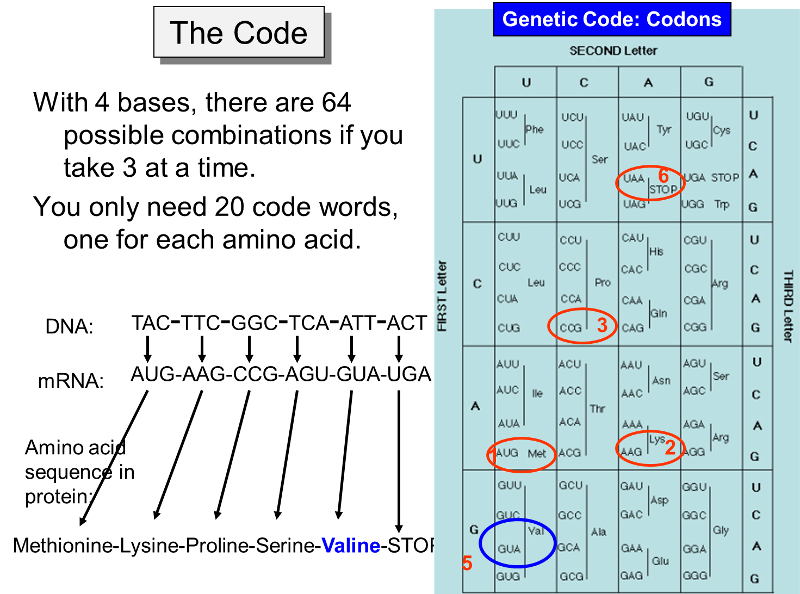
The figure above makes it clear that the order of the codons within a gene (a segment of DNA encoding for a specific protein) specifies the amino acid sequence in the protein. The start signal for protein synthesis is the codon AUG, which specifies incorporation of the amino acid methionine. When the mRNA attaches to a ribosome, enzymes look for the AUG codon, not only as a start signal, but also as a means of knowing exactly which is the first letter of each of the three-letter codons. For example, a messenger RNA might have the sequence of codons shown in the illustration above, i.e.,
AUG-AAG-CCG-AGU-GUA-UGA-... etc.
However, if the signal to start was shifted by one nucleotide (e.g., starting at the first "U" instead of the "A"), the codons would be read as:
UGA-AGC-CGA-GUG-UAU- ..etc.
and this would result in synthesis of an very different sequence of amino acids. Errors in the sequence of amino acids can, in fact, result from mutations, as described below.
The video below gives a detailed summary of the events that occur during translation of the mRNA template to a protein.
This next video is an excellent illustration of transcription and translation, but it illustrates these in a way that provides a real-time approximation.
|
An Interesting Variation by HIV
The human immunodeficiency virus (HIV) is known as a retrovirus. It consists of a single strand (molecule) of RNA inside a protein coat. When HIV binds to a T lymphocyte it enters the lymphocyte and sheds its protein coat. A viral enzyme called reverse trancriptase then uses the strand of viral RNA as a template to create a molecule of DNA which can become incorporated into the DNA of the infected host cell. In this case, RNA is being used to create a molecule of DNA, and the process has been dubbed "reverse transcription."
|
Most of the DNA in eukaryotic cells is contained in the chromosomes within the membrane-bound nucleus, but the mitochondria also have small amounts of DNA (mitochrondrial DNA or mtDNA). As you will recall, mitochondria are membranous subcellular organelles within which there are chains of enzymes that generate cellular energy in the form of ATP (adenosine triphosphate) through a process called oxidative phosphorylation. In addition to their role in production of ATP, mitochondria also regulate apoptosis (programmed cell death) [for more on apoptosis see the section on Apoptosis in the module on the Biology of Cancer]. Mitochondria also play a role in the synthesis of cholesterol and heme (a component of hemoglobin, the oxygen carrying molecules in red blood cells). Mitochondrial DNA consists of 37 genes. Thirteen of these provide the genetic code for synthesizing the enzymes involved in oxidative phosphorylation, and the rest encode the transfer RNAs (tRNA) and ribosomal RNA (rRNA) required for synthesis of the enzymes for oxidative phosphorylation.
Inherited mutations in mitochondrial DNA can also cause a variety of problems with growth, development, and function throughput the body as a result of impaired ability to generate ATP. These conditions can produce muscle weakness and wasting, diabetes, kidney failure, heart disease, dementia, hearing loss, visual problems. In addition, mitochondria can also undergo somatic mutations (non-inherited) which may contribute to aging and age-related diseases.
Additional Resources for Mitochondrial DNA
For more information about conditions caused by mitochondrial DNA mutations:
The preceding pages describe how the genetic code in DNA specifies the assembly of proteins, which are polypeptides, meaning that their primary structure consists of a long linear assembly of amino acids. As the amino acids are being linked together into a growing polypeptide begins to fold as a result of the interactions among the amino acids. Different segments of a protein can assume different shapes: random coils, helices, and zig-zagging segments that form sheet-like structures; these secondary structures are then folded in ways that establish the tertiary structure, and these fold yet again to create the final quaternary structure, as shown in the graphic below.
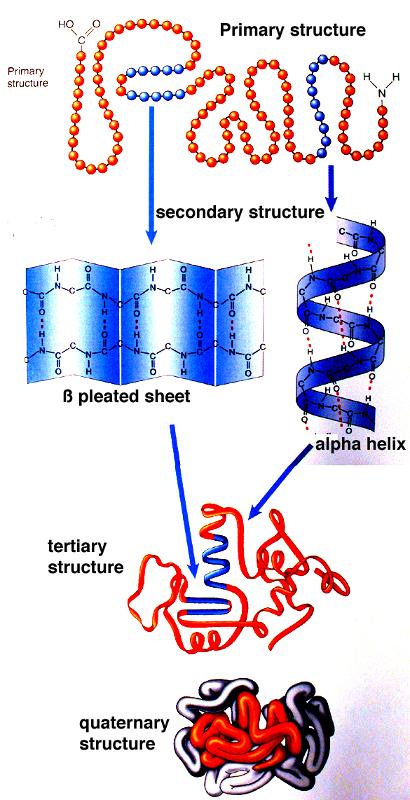
Source: http://academic.brooklyn.cuny.edu/biology/bio4fv/page/3d_prot.htm
Each protein has a specific shape that determines its function. The illustration below summarizes just some of the many functions carried out by proteins.
There are many thousands of enzymes that participate in synthesis and metabolism.
All humans have the same set of genes, and the sequence of our base pairs is remarkably similar. However, this doesn't mean that we all have exactly the same nucleotide sequence in our genome. If this were the case, then all humans would be clones having exactly the same genetic code. The passage of the genetic code from generation to generation via the sperm and ova of our ancestors requires replication of DNA, and while replication is remarkably precise, errors occasionally occur and produce changes in the base sequence. It is these changes that produce the variation that makes each of us genetically unique (except for identical twins), and they also drive evolution of species
Mutations are random changes in the sequence of base pairs in DNA, and mutagens are factors that cause mutations, e.g., chemicals or radiation (UV light, x-rays, gamma radiation). Mutagens result four patterns of alteration in the base sequence:
The effects of these alterations depend on several factors.
Substitution of one base pair for another produces the smallest change in the code, but it's effects can range from none to massive depending on the details.
The example below shows an initial sequence of base pairs on the left, the mRNA transcript, and the sequence of amino acids that would result. The right portion shows that substitution of thymine (T) for adenine (A) changes the third base in the mRNA transcript, but the amino acid sequence is unchanged, because GUU and GUA are both codons for the amino acid valine. In fact, there are four codons that specify insertion of the amino acid valine: GUU, GUC, GUA, and GUG. If a normal gene had a GUU codon (specifying insertion of valine) at a particular place, changing the third base from uracil (U) to adenine (A) would still produce a protein that had the amino acid valine in the correct position. Consequently, the protein would be unchanged, and the mutation would be "silent."
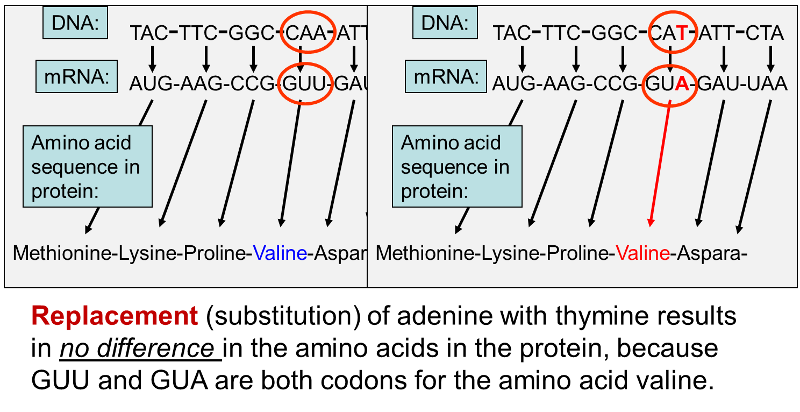
In other cases, however, substitution of a single base pair leads to a change in one of the amino acids in the protein product as shown below.
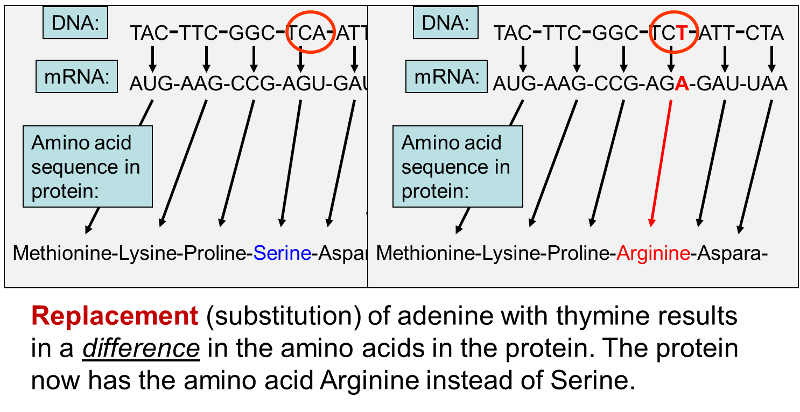
Once again, an adenine (A) in DNA has been replaced by a thymine (T), but in this case the codon in the mRNA changes from AGU, which is a codon for the amino acid serine to AGA , which codes for the amino acid arginine. Changing one amino acid in a protein may have no discernible effect on the protein's function, but at times, changing even a single amino acid can have a massive effect. An example of this is seen with sickle cell anemia.
Sickle cell anemia is due to an inherited mutation in the gene that encodes for the β- chain of hemoglobin. Hemoglobin is a protein found in red blood cells which plays a key role in transporting oxygen from the lungs. It is composed of four protein subunits: two α- and two β-chains. In sickle cell anemia the key defect is caused by a mutation that replaces the hydrophilic amino acid glutamic acid (glutamate) with the hydrophobic amino acid valine at the sixth position of the β-chains. This causes the hemoglobin molecules to stick together when oxygen levels are low. As a result, the hemoglobin molecules form long fiber-like chains that distort the shaped of the red blood cells. Red blood cells are normally biconcave disks that flow easily through arteries and capillaries, but sickling causes red blood cells to stick to one another and occlude small arteries causing ischemia in various tissues and organs.
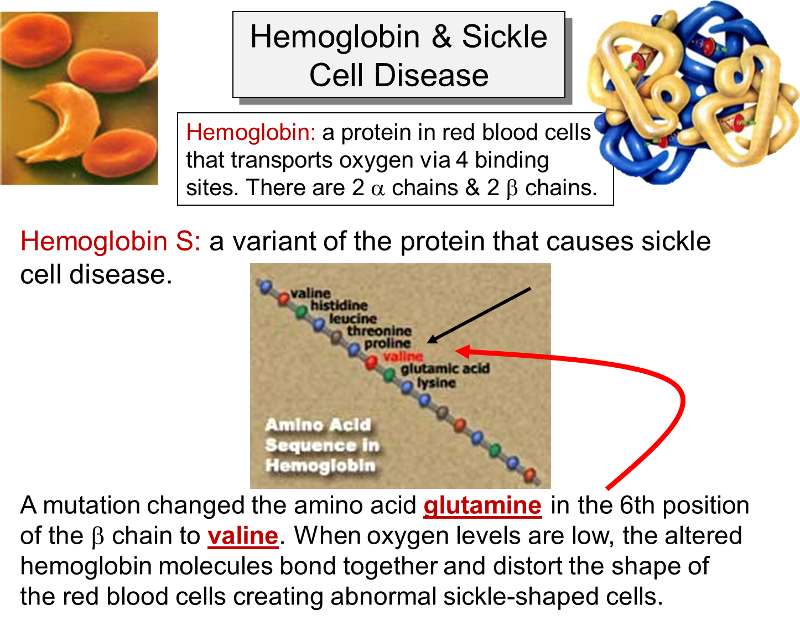
Link to more information on sickle cell disease.
In contrast to substitution of a single base, addition or deletion of a single base cause a substantial disruption of the sequence of amino acids in the protein product. This is because the mRNA transcript is read as three-letter codons, and insertion or deletion of a single base causes a frame shift in the sequence that throws off all of the downstream codons. Consider the following hypothetical sequence of DNA and the mRNA transcript it would produce and the final amino acid sequence. For illustration, I begin with triplets with the same base, e.g., AAA, TTT, etc.
DNA -- AAA-CCC-TTT-GGG-GGG- TTT-CCC-TTT-AAA-AAA - etc.
mRNA -- UUU-GGG-AAA-CCC-CCC-AAA-GGG-AAA-UUU-UUU- etc.
Amino Acids -- Phe -- Gly-- Lys--Pro--Pro --Lys --Gly --Lys--Phe--Phe--etc.
Insertion of a single base might result in the following:
DNA -- AAA-ACC-CTT-TGG-GGG-GTT-TCC-CTT-TAA-AAA-A etc.
mRNA --UUU-UGG-GAA-ACC-CCC-CAA-AGG-GAA-AUU-UUU-U- etc.
Amino Acids -- Phe--Trp- Glu--Thr--Pro -Gln -Arg-Glu--Ile--Phe--etc.
The insertion of a base advances all of the letters by one position, but the codons are still read as triplets, so the code is thrown off from the point of the insertion, and most of the amino acids are changed, although occasionally the original amino acid is retained by chance such as the Pro (proline) and the terminal Phe (phenylalanine) in the example above.
Deletion of a single base has the same effect, i.e., it causes a frame shift that changes all of the codons downstream from the point of the deletion.
The short video below illustrates the effects of substitution, insertion, and deletion of single base pairs.
The other mechanism that has been described for mutation is the relocation of an entire segment of nucleotides. This is illustrated below starting with the same hypothetical sequence of bases in DNA.
DNA -- AAA-CCC-TTT-GGG-GGG- TTT-CCC-TTT-AAA-AAA - etc.
mRNA -- UUU-GGG-AAA-CCC-CCC-AAA-GGG-AAA-UUU-UUU- etc.
Amino Acids -- Phe -- Gly-- Lys--Pro--Pro --Lys --Gly --Lys--Phe--Phe--etc.
Relocation of a segment of bases might result in:
DNA -- AAA-CCT- CCC-TTT-AAC-TTT-GGG-GGG-TTA-AAA - etc.
mRNA -- UUU-GGA-GGG-AAA-UUG-AAA-CCC - CCC - AAU-UUU- etc.
Amino Acids -- Phe -- Gly-- Lys--Pro--Leu --Phe --Pro -- Pro--Asn--Phe--etc.
This kind of relocation can have variable effects depending on the size of the relocated segment and whether or not it also cause frame shift errors.
We noted earlier that an error in replication of DNA can result in substitution of one base for another. For example, the human genome might have the following segment of base pairs in double-stranded DNA:
...CGATATTCCTATCGAATGTC....
...GCTATAAGGATAGCTTACAG...
However, a substitution of a single base pair (changing the bold-faced TA pair to GC) might result in a sequence that read as follows:
...CGATAGTCCTATCGAATGTC....
...GCTATCAGGATAGCTTACAG...
We saw earlier that if a substitution like this occurred within a gene, i.e., within a coding area, it could cause a disease like sickle cell anemia, but it is also possible that it might have no discernible effect. Remember, however that coding areas (genes) occupy only 3-5% of our DNA, and substitutions like occur more commonly in the much more extensive non-coding areas between the genes. The accumulation of these random substitutions over time has resulted in many small differences in the sequence of base pairs from person to person. These small differences are found only about once every 300 base pairs on average. However, since we have 3 billion base pairs, this means that there are about 10 million of these small differences in our genome. These small variations have been dubbed "single nucleotide polymorphisms" or SNPs (pronounced "snips"). Each SNP is a difference in a single base in DNA.
In the discussion of mutations we saw that substitution of a single base (a SNP) within a coding area may or may not have an effect on phenotype. In contrast, SNPs occurring in non-coding areas of the genome have no effect on phenotype (characteristics). Nevertheless, each person has a unique SNP pattern, however, and this is potentially useful in a number of ways.
For more information about SNPs go to the two following links:
The sequence of bases in the human genome is remarkably similar from person to person, but over hundreds of thousands of years of evolution SNPs and other mutations have been introduced into the human gene pool. Some of these mutations produce alterations in gene products that are fatal, and these mutations are extinguished. However, other mutations in germ cells (sperm and eggs) can be passed along from generation to generation, and they provide the basis for the many variations in phenotype that make each of us unique. Over time, mutations have created variants of genes that are responsible for differences in the color of our hair, our eyes, and our skin. Mutations influence our intelligence, our height, our weight, our personalities, our blood pressure, our cholesterol levels, and how fast we can run. Mutations have introduced gene variants that encode for slightly different proteins, which in turn, influence all aspects of our phenotype. It is important to emphasize an individual's phenotype is not solely the result of their genome; instead, phenotype is the result of the interaction between and individual's genome and their environment from the time of conception until death.
When SNPs and other mutations create variants or alternate types of a particular gene, the alternative gene forms are referred to as alleles. In other words, a given gene can have multiple alleles (i.e., alternate forms). Some genes have just a few alleles, but others have many.
Recall also that chromosomes come in pairs. Humans have 22 pairs of autosomal chromosomes with the same gene in both members of a given pair) and one pair of sex chromosomes, which are designated XX in females and XY in males. The X and Y chromosomes are physically different from one another in that the Y chromosome is much shorter, and the Y chromosome only has about nine gene loci that match those on the X chromosome. This means that, except for the genes on an XY pair of chromosomes, we have two copies of each gene - one from each of our parents. The alleles that we receive from each parent might be the same (homozygous) or they might differ (heterozygous). The figure below schematically depicts a pair of chromosomes and shows three hypothetical genes: hair color, body height, and multiple lipoma formation.
Since there are two copies of each gene, there are two alleles, which may be the same or different. The figure below shows a hypothetical example in which there is an allele for red hair on one chromosome and an allele for brown hair on the other.
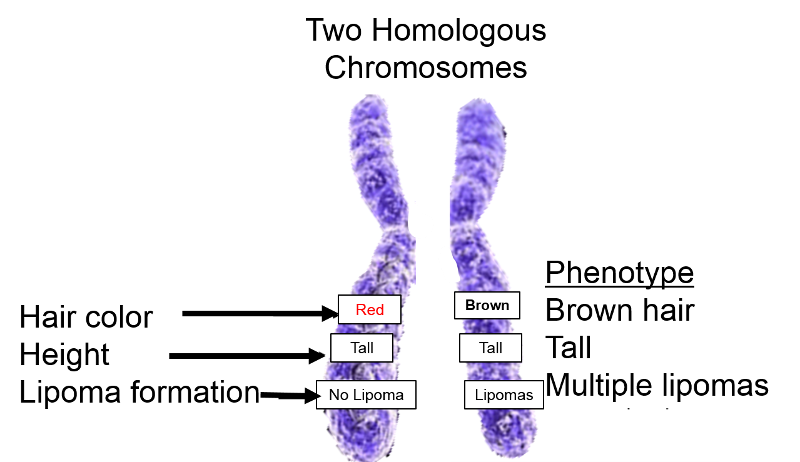
(Note that there may be many alleles for some genes, but normally we each have two alleles for each gene on our autosomes. Note also that in the hypothetical illustration to the right the alleles for the multiple lipoma trait are also different.
The obvious question that arises is, what happens when the two alleles that are present differ? What will the phenotype be? The answer depends on whether one allele is dominant over the other.
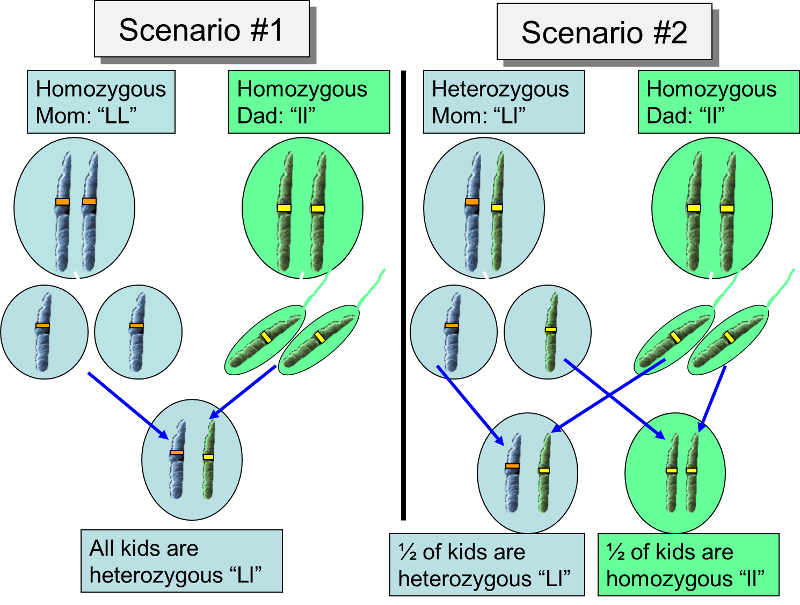
A dominant allele is one that is expressed to a greater degree than the other allele that is present. For example, one possible scenario for the differing lipoma alleles is shown below.
What about another scenario in which the mom is heterozygous and the dad is homozygous recessive?
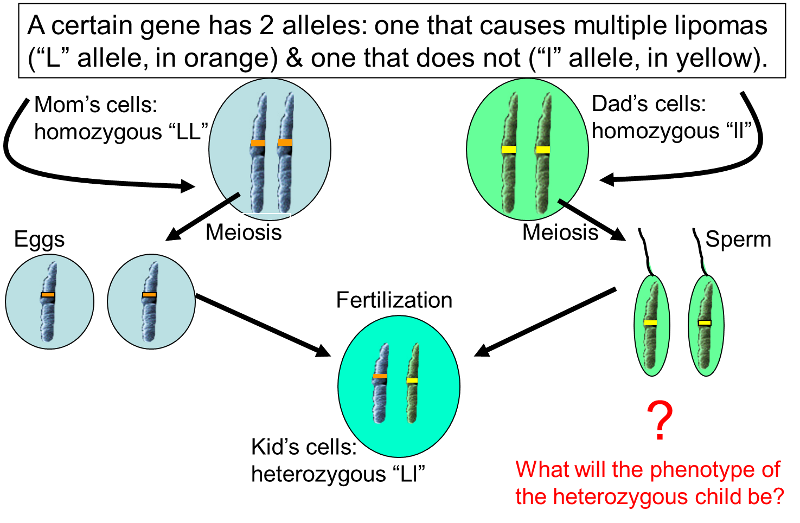
Mom is homozygous for the multiple lipoma trait (designated as "LL"), while Dad is homozygous for the absence of lipomas (designated "ll"). Mom can only contribute an "L" allele to her offspring, and Dad can only contribute the "l" allele, so all of their children will be heterozygous ("Ll"). In this particular case, heterozygous "Ll" individuals will all have multiple lipomas, because the multiple lipoma allele is dominant, while the alternate "l" allele is recessive.
For some alleles there is no dominance, and phenotype results from both alleles being expressed or from a blending of phenotype. The expression is an "average" or combination of the two traits.
Example: Major blood type in humans.
In humans, for example, there is a specific gene that codes for the protein that determines an individual's major blood type, which can be A, B, AB, or O. This is determined by a single gene that has three alleles that can code for:
While there are three alleles, each of us has just two of them, so the possible combinations and the resulting blood types are those shown in the table below.
|
Alleles |
Phenotype (Blood Type) |
|---|---|
|
AA |
A |
|
AO |
A |
|
BB |
B |
|
BO |
B |
|
OO |
O |
|
AB |
AB |
The 3 billion base pairs in the human genome are remarkably similar from one person to another, but over tens of thousands of years random mutations in genes have introduced many variants. We saw previously that mutations can result in
Depending on the function of the gene and the magnitude of change, a mutation may or may not be compatible with life. A fetus with a critical mutation may be unable to survive, leading to a miscarriage (spontaneous abortion). Non-fatal mutations can result in protein alterations that alter characteristics like hair or eye color, or they can produce proteins that function better or worse than the usual protein. All of these accumulated differences in genes are what distinguish one person from another and make each of us unique. (Identical twins are born with exactly the same genome, but a host of environmental and epigenetic factors produce differences even in identical twins.)
These occasional random mutations are responsible for driving the evolution of species by sometimes conferring a survival advantage. Consider two brief examples:
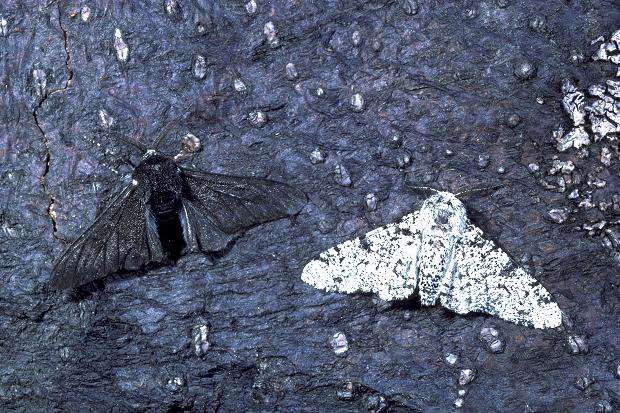

Source: http://www.openwindowlearning.com/quizzes/the-role-of-genetic-diversity-in-evolution/
Charles Darwin postulated that if a mutation confers some advantage (e.g., resistance to penicillin, or better camouflage, or faster muscles, etc.) than those organisms will be better able to compete in an environment with limited resources or other environmental pressures (such as penicillin or predatory moths). These "fitter" organisms will therefore have more opportunity to thrive and reproduce, and their numbers will increase. As a result, the frequency of the mutant gene will increase in the organisms in the particular environment that gives those with the mutation an advantage.
Having an allele for the sickle cell form of hemoglobin would seem to be a bad thing, but that depends on both the genotype and the environment. The gene for sickle cell disease follows a Mendelian pattern of inheritance (described briefly on page 11). The allele for sickle cell hemoglobin is designated HbS, and there are several genotypes that are possible:
Those who are homozygous for HbS have significant health problems and poor outcomes, but those who are heterozygous only have clinical manifestations under certain circumstances when the oxygen concentrations in blood dip, e.g., in high altitudes or with heavy physical activity. Without these stresses, heterozygous persons function normally. In fact, heterozygous HbA-HbS individuals have an advantage in locations where malaria is endemic, because their red blood cells are fragile and tend to lyse (fall apart), when infected with malaria. As a result, heterozygous individuals have a survival advantage in areas where malaria is endemic, and given this survival advantage the sickle cell allele tends to persist in environment where malaria is endemic as demonstrated by the graphic below which shows the prevalence of the sickle cell allele in the top panel and prevalence of malaria below.
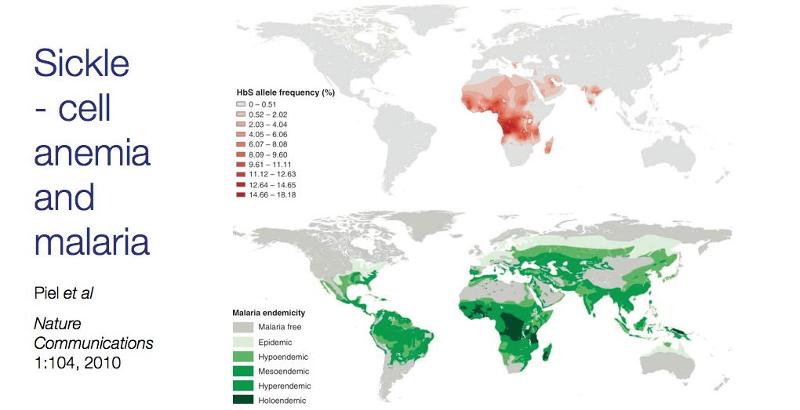
Source: Piel et al.: Nature Communications1:104, 2010
By gene expression we mean the transcription of a gene into mRNA and its subsequent translation into protein. Gene expression is primarily controlled at the level of transcription, largely as a result of binding of proteins to specific sites on DNA. In 1965 Francois Jacob, Jacques Monod, and Andre Lwoff shared the Nobel prize in medicine for their work supporting the idea that control of enzyme levels in cells is regulated by transcription of DNA. occurs through regulation of transcription, which can be either induced or repressed. These researchers proposed that production of the enzyme is controlled by an "operon," which consists a series of related genes on the chromosome consisting of an operator, a promoter, a regulator gene, and structural genes.
The operator gene is the sequence of non-transcribable DNA that is the repressor binding site. There is also a regulator gene, which codes for the synthesis of a repressor molecule hat binds to the operator

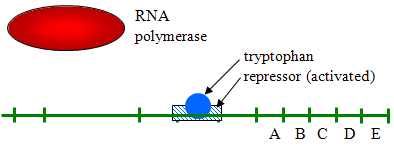
Source: http://biowiki.ucdavis.edu/Under_Construction/BioStuff/BIO_101/Reading_and_Lecture_Notes/Control_of_Gene_Expression_in_Prokaryotes
Eukaryotic cells have similar mechanisms for control of gene expression, but they are more complex. Consider, for example, that prokaryotic cells of a given species are all the same, but most eukaryotes are multicellular organisms with many cell types, so control of gene expression is much more complicated. Not surprisingly, gene expression in eukaryotic cells is controlled by a number of complex processes which are summarized by the following list.
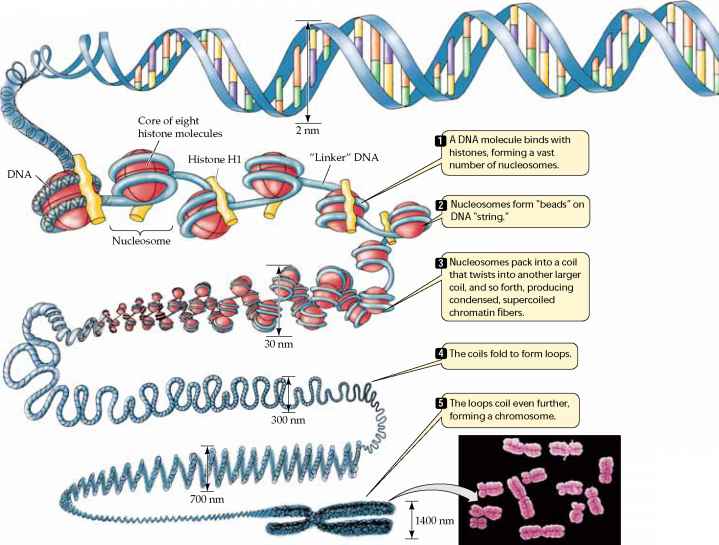
Source: http://www.78stepshealth.us/plasma-membrane/eukaryotic-chromosomes.html
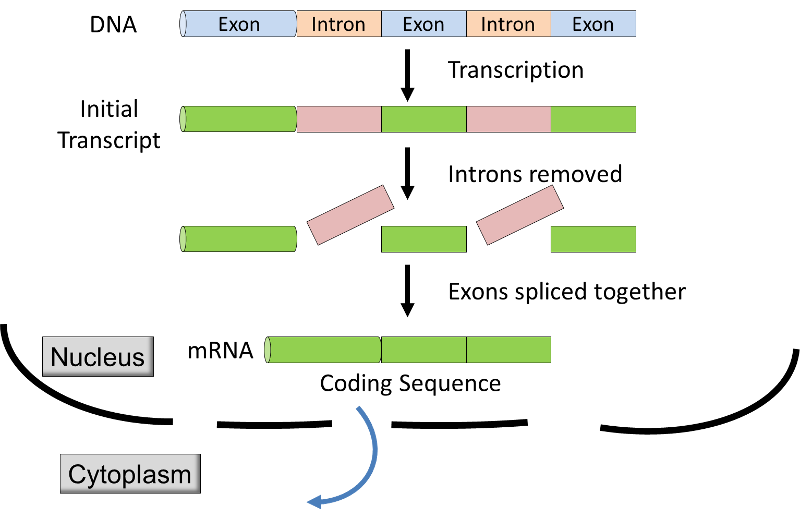
Source: http://unmug.com/category/biology/organisation-control-of-genome/
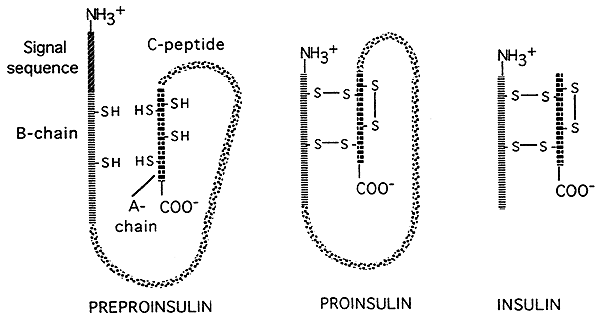
Source: http://www.nbs.csudh.edu/chemistry/faculty/nsturm/CHE450/19_InsulinGlucagon.htm
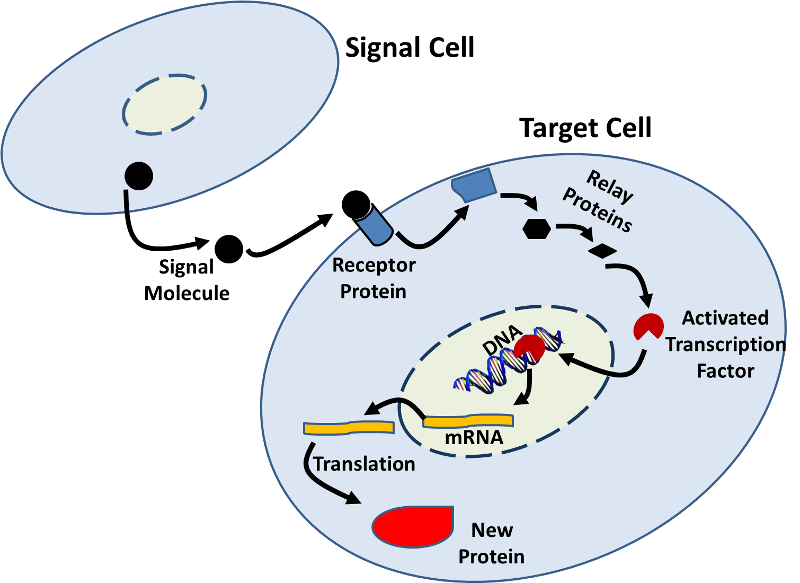
Source: http://sites.saschina.org/emily01px2016/2014/11/23/a-variety-of-intercellular-and-intracellular-signal-transmissions-mediate-gene-expression/
Some RNA virus will invade cells and introduce double-stranded RNA which will use the cells machinery to make new copies of viral RNA and viral proteins. The cell's RNA interference system (RNAi) can prevent the viral RNA from replicating. First, an enzyme nicknamed "Dicer" chops any double-stranded RNA it finds into pieces that are about 22 nucleotides long. Next, protein complexes called RISC (RNA-induced Silencing Complex) bind to the fragments of double-stranded RNA, winds it, and then releases one of the strands, while retaining the other. The RISC-RNA complex will then bind to any other viral RNA with nucleotide sequences matching those on the RNA attached to the complex. This binding blocks translation of viral proteins at least partially, if not completely. The RNAi system could potentially be used to develop treatments for defective genes that cause disease. The treatment would involve making a double-stranded RNA from the diseased gene and introducing it into cells to silence the expression of that gene. For an illustrated explanation of RNAi, see the short, interactive Flash module at http://www.pbs.org/wgbh/nova/body/rnai-explained.html
The RNA interference system is also explained more completely in the video below from Nature Video.
Our genome is established when fertilization takes place, and the code remains unchanged throughout our life, except for mutations that may occur in individual cells. Nevertheless, the previous page outlined many internal mechanisms that operate to control the expression of specific genes. In addition, we now know that many external factors (epigenetics) can affect the timing of the gene expression, the degree of expression, and the eventual phenotype that is expressed. These external factors can produce small modifications to DNA, such as addition of metal ions, addition or removal of acetyl groups or methyl groups to DNA or to the histones that control the wrapping and packing of DNA. Attachment of methyl groups appears to reduce transcription or even shut it off; attachment of acetyl groups to histones turns genes on. These biological changes to the genome is known as 'epigenetic factors', i.e., changes occurring above the level of the genome.
The methyl group
![]()
The acetyl group
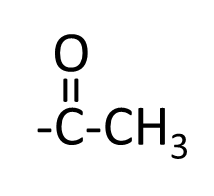
Modification of DNA by the methyl group:
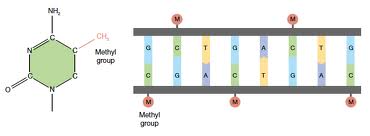
In essence, the DNA in our cells provide the code for making functional proteins, and the epigenetic factors act as switches which turn genes off and on. Epigenetic factors are likely to play many important roles, such as:
This is an 18:40 min video from TEDxOU by Courtney Griffin that provides an excellent explanation of the interaction among:
In summary, all of our traits and characteristics (our phenotype) are the result of an interaction between our genome (all of the genes we inherit) and environmental factors. Some environmental factors (including our diet, our behaviors, and a myriad of environmental exposures) influence our phenotype through non-genetic mechanisms. For example, one might have a number of genes that predispose an individual to being lean; however, such an individual might still become overweight or obese despite their "lean genes" as a result of chronically overeating. Yet other epigenetic factors from the environment can modify the genome in subtle ways without actually changing the code.
For more information on epigenetics explore the following web site: http://learn.genetics.utah.edu/content/epigenetics/
Did you know?
Pink hydrangeas can be made to turn blue by adding aluminum sulfate to the soil.

Prokaryotes reproduce by the relatively simple process of binary fission. The single chromosome replicates and each copy attaches to a different location on the cell membrane. The cell membrane then begins to invaginate and eventually separates into two genetically identical bacteria. A similar process is used to replicate mitochondria within eukaryotic cells, but the overall process of cell replication in eukaryotes is more complicated (see below).
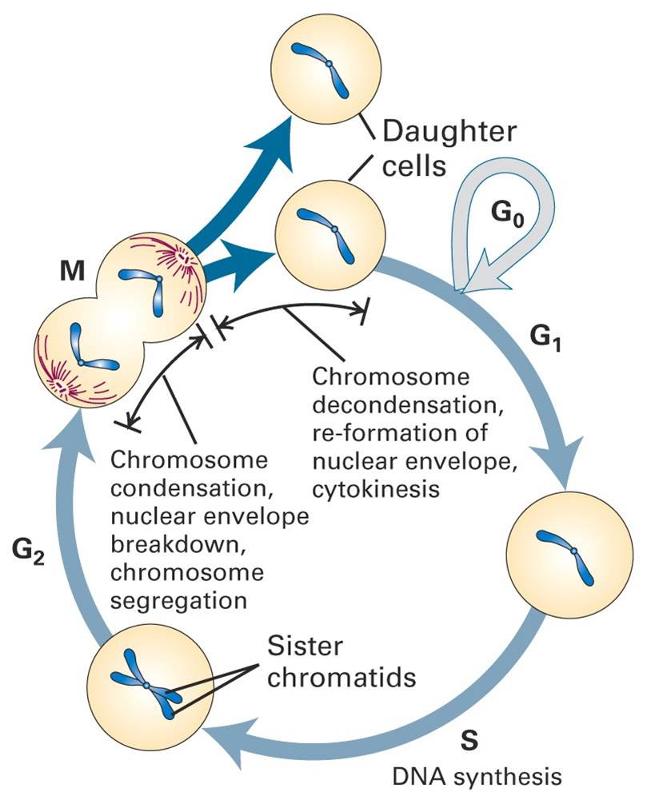
Adapted from http://www.slideshare.net/SurenderRawat3/cell-cycle-checkpoints-apoptosis-and-cancer
Mitosis is the process by which eukaryotic cells replicate by dividing into two genetically identical cells. It is the process by which new cells are formed in the growing embryo and after birth, and mitosis also replaces cells that have died or been shed. In humans some cells retain the capacity to divide throughout life. These "stem cells" divide by mitosis and produce daughter cells which then differentiate into a particular cell type. This provides a way of replacing cells, such as skin cells; the epithelial cells that line the respiratory, digestive, and urogenital tracts; and blood cells. Benign and malignant tumors also growth through mitosis.
Cells normally follow a carefully controlled cell cycle, depicted below.
Many of our cells are mature functioning cells that are not actively dividing. These are cells in the G0 phase; this is sometimes called the "resting phase," but these cells are actively functioning, and they are resting only in the sense that they aren't replicating. The phases in dividing cells are as follows:
The cell cycle is normally carefully controlled by a number of biochemical mechanisms. Loss of control mechanisms can result in abnormal cell division and a progression to to tumor formation. This is discussed in greater detail in the online module on cancer.
Meiosis is the specialized process by which gametes (sperm and eggs) are produced for sexual reproduction in the ovaries and testes. Recall that humans have 22 pairs of homologous chromosomes and one pair of sex chromosomes; one member of each pair came from the mother, and the other from the father. The 46 chromosomes are referred to as the diploid (2n) number, because there are two of each. In order for the fertilized egg to end up with the correct diploid number, sperm cells and eggs must be produced such that each has only one chromosome from each pair. In other words, gametes have only 23 chromosomes (referred to as the haploid number (1n). Meiosis, then, is the process by which specialized diploid stem cells in the ovary (oogonia) and testes (spermatogonia) produce eggs and sperm which have a haploid number of chromosomes. Thus, each gamete has 23 chromosomes (one from each of the 22 homologous pairs + 1 sex chromosome).
The video below shows the differences between mitosis and meiosis.
Meiosis produces sperm and eggs with novel mixtures of the original parental chromosomes due to:
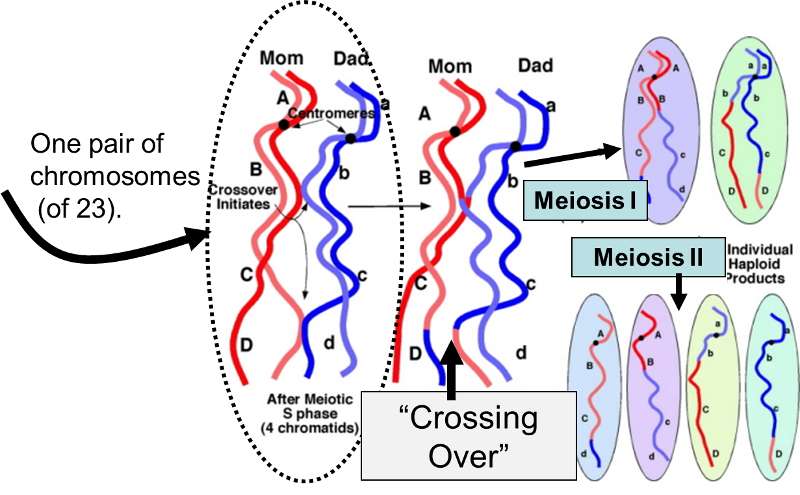
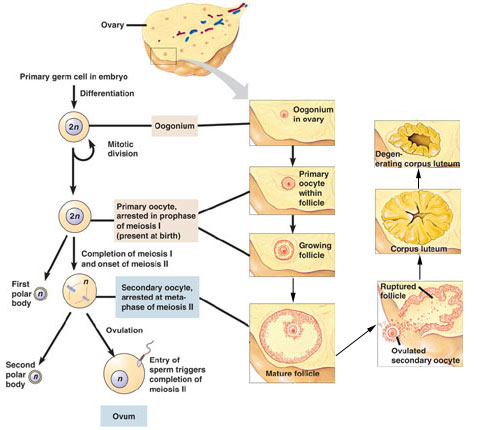
Source: http://biologyandmedicineanimation.blogspot.com/2014/12/oogenesis.html
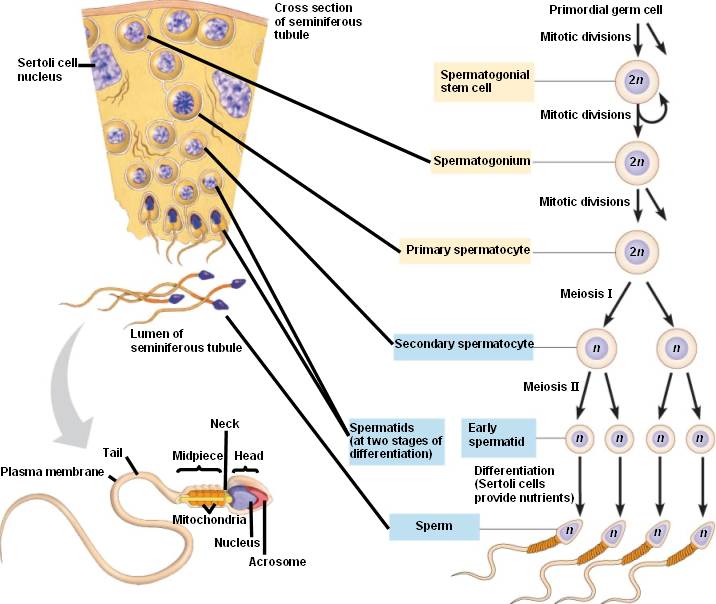
Source: http://bio1152.nicerweb.com/Locked/media/ch46/spermatogenesis.html
The diagram below summarizes the processes of meiosis, fertilzation, and mitosis. Note that the numbers indicate either the diploid number of chromosomes in cells (46) or the haploid number (23) in gametes.
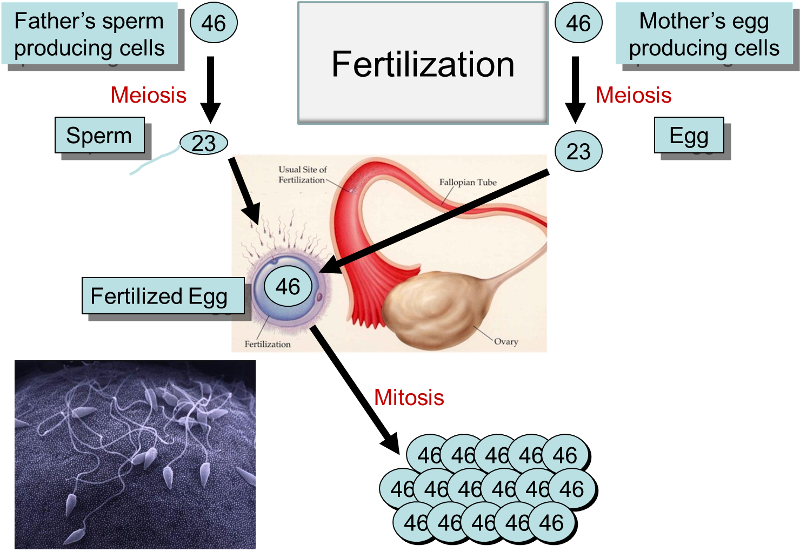
Gregor Mendel was an Austrian monk who formulated some of the fundamental principles regarding the inheritance of traits. Between 1856 and 1863 he performed thousands of experiments in which he cross-bred pea plants with dichotomous characteristics such as color (e.g., yellow or green). Several conclusions were drawn from his studies:
The gene that determines whether multiple lipomas will form (referred to on page 6) illustrates a Mendelian pattern of inheritance.

In this case, the "L" allele that encodes for multiple lipomas is dominant over the "l" allele which does not cause lipomas.

With a dichotomous trait like this one can one predictions about the proportions of offspring by using a Punnett square which shows the four possible pairs of alleles that can occur in the offspring. In Scenario #1 above, the Punnett square demonstrates that only heterozygous gene pairs are possible, so all of the offspring will have multiple lipomas, since the lipoma allele is dominant.
Table 1: Punnett Square for Offspring of a Homozygous Dominant (LL) Mother and a Homozygous Recessive (ll) Father
|
|
|
Father's Alleles |
|
|
|
|
l |
l |
|
Mother's Alleles |
L |
Ll |
Ll |
|
L |
Ll |
Ll |
|
All of the children wil he hetorzygous (Ll) and have the dominant trait.
Table 2: Punnett Square for Offspring of a Heteroygous (Ll) Mother and a Homozygous Recessive (ll) Father
|
|
|
Father's Alleles |
|
|
|
|
l |
l |
|
Mother's Alleles |
L |
Ll |
Ll |
|
l |
ll |
ll |
|
In this case half of the offspring (on average) will be heterozygous and have multiple lipomas, and the other half will be homozygous recessive and be free of lipomas.
Table 3: Punnett Square for Gender
|
|
|
Father's Sex Chromosomes |
|
|
|
|
X |
Y |
|
Mother's Sex Chromosomes |
X |
XX |
XY |
|
X |
XX |
XY |
|
The mother has XX sex chromosomes and the father has XY, so half of the offspring are predicted to be female, and half will be male.
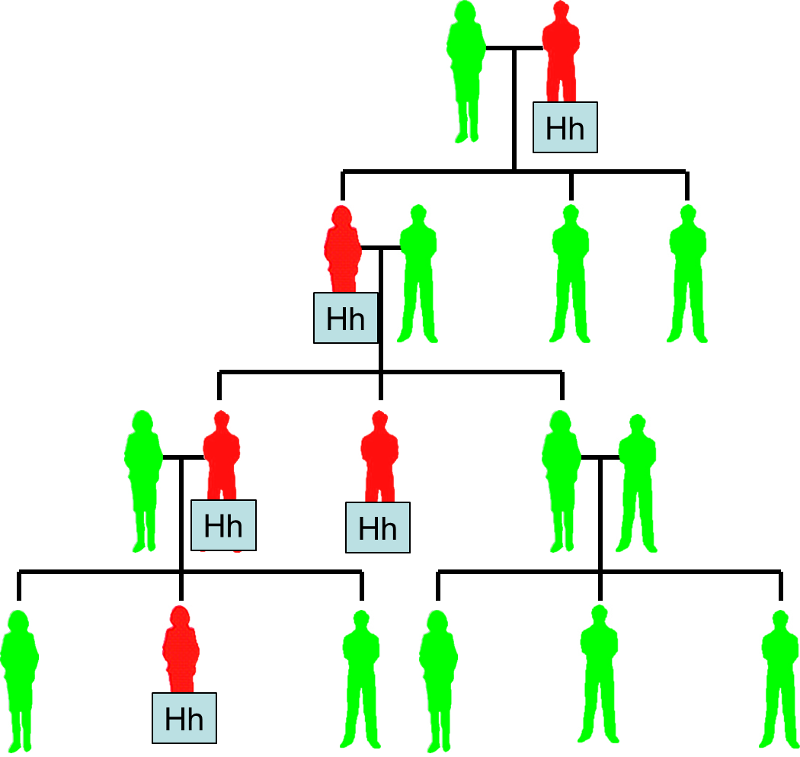
Some disease are inherited, and the pattern of appearance within a family tree will depend on whether the faulty allele is dominant or recessive compared to the normal allele. For example, the allele for Huntington's disease is dominant. If a heterozygous (Hh) man with Huntington's disease and a normal woman (hh) have children, some of them (about half on average) will have the disease (individuals shown in red). With a dominant allele like this, the disease occurs fairly frequently in the family tree.
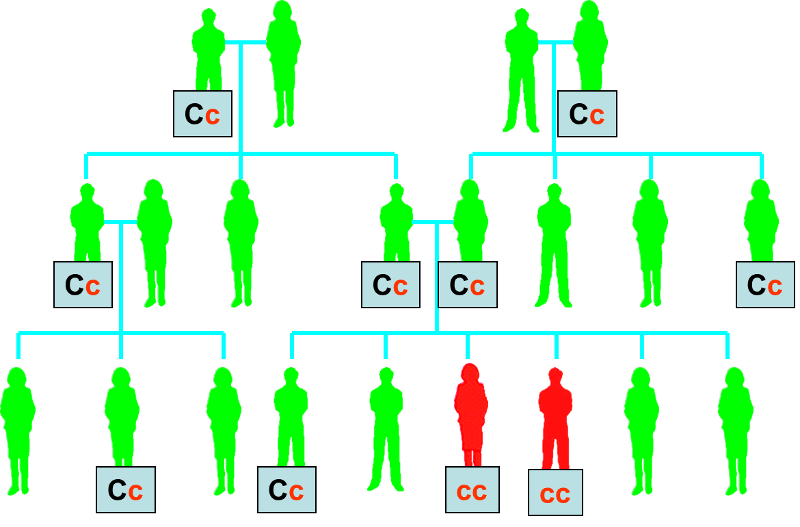
In contrast to Huntington's disease, cystic fibrosis is caused by a recessive allele, meaning that individuals who are heterozygous for the cystic fibrosis allele (shown as Cc below) will not manifest any signs or symptoms of cystic fibrosis. As a result, the cystic fibrosis allele can be passed along a family tree with only sporadic appearance of individuals who have signs and symptoms of cystic fibrosis because they are homozygous for the recessive allele (cc).
Huntington's Disease
Cystic Fibrosis
Mendel's studies focused on dichotomous traits in plants, such as the color of peas (green or yellow) and plant size (tall or dwarf), but many traits have continuous distributions, such as height, weight, and intelligence. Galton was a contemporary of Mendel's who studied the inheritance of continuous characteristics. The idea that characteristics might be blended or averaged occurred to him when he noted that very tall fathers tended to have sons shorter than themselves, and extremely short fathers tended to have sons taller than themselves. He referred to this as "regression to mediocrity," and he concluded that height doesn't follow the inheritance patterns of the dichotomous traits that Mendel studied and that the phenomenon of dominance didn't apply here.
Mendelian inheritance patterns predicted some diseases, but only a few, and Galtonian genetics was limited by the inability to predict outcomes. R. A. Fisher, a British statistician and evolutionary biologist, was able to reconcile these two patterns of inheritance by showing that the inheritance of quantitative traits can be reduced to Mendelian inheritance if multiple genes are involved. For example, suppose the average height in a population is 68", and height is determined by one gene with 3 possible alleles: H0 (which neither adds nor subtracts from the average); H+2 (which adds 2" to height), and H-2 (which subtracts 2" from height). Suppose also the the H0 allele is twice as common as the other two alleles in the population. If these are co-dominant alleles, a Punnett square would predict the following inheritance patterns distribution of heights:
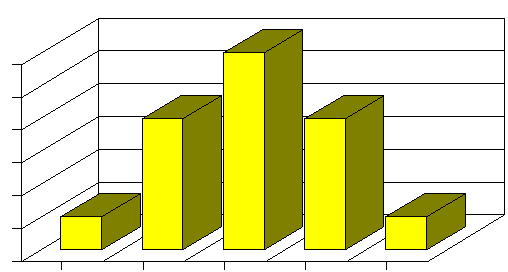
Adapted from http://www.uic.edu/classes/bms/bms655/lesson11.html
Now, if instead of just one gene, there were two genes that determined height, and both of them had three possible alleles as described above, then a Punnett square would predict a distribution with more categories and finerdifference among the categories.
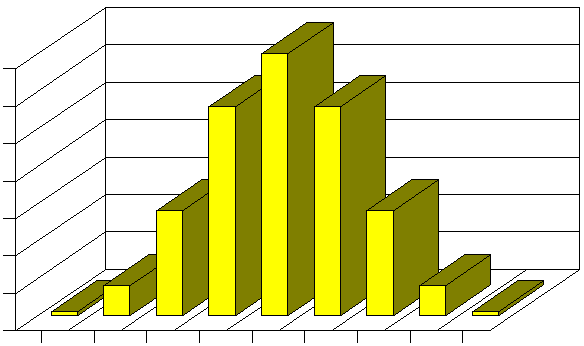
Adapted from http://www.uic.edu/classes/bms/bms655/lesson11.html
It is easy to imagine that if there were three or more genes that were also determinants of height, the distribution would increasingly conform to a Gaussian distribution. In fact, a similar model with three gene loci, each with three alleles looks very much like a bell-shaped distribution. As a result, continuously distributed characteristics are likely to be determined by a finite number of genes which have co-dominant alleles. And, once again, it is important to point out that many environmental factors are likely to interact with the genotype to produce the final phenotype.
Earlier in this modules it was noted that X and Y chromosomes are physically different from one another in that the Y chromosome is much shorter, and the Y chromosome only has about nine gene loci that match those on the X chromosome. As a result, almost all of the alleles on a male's single X chromosome are expressed, since there is no alternative dominant allele to mask them. This results in a distinct inheritance pattern for traits that are encoded on the X chromosome. For example, there are many types of color blindness. All of these conditions are inherited, and most of them are "sex-linked" because they are caused by defective alleles carried on the X chromosome. Red-green color blindness is a fairly common, mild form of color blindness which can be found in about 6% of the male population; it is far less common in females. This form of color blindness is caused by a recessive allele, and the inheritance pattern is illustrated in the figure below.
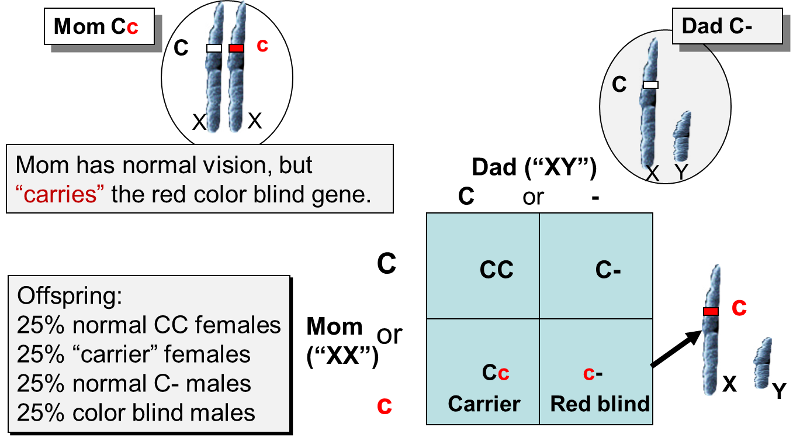
The defective allele (c) is only carried on the X chromosome, as is the normal allele (C), which is dominant. A heterozygous female, as shown above, would have normal color vision, but she would be a "carrier" of the allele for red-green color blindness. Now suppose that she marries a man with normal red-green color vision (he only has the C allele), and they have children. On average, half of the offspring will inherit mom's defective X chromosome, and half will inherit the X chromosome with the normal allele. Dad can contribute either an X chromosome with a normal allele or a Y chromosome with no allele for color vision. The Punnett square illustrates the possible combinations of alleles that will occur in the offspring. As you can see, there are four possible results. Both males and females can inherit the allele for color blindness, but it will not be expressed in any of the females, because the normal allele is dominant, so the heterozygous "Cc" females will be carriers of the trait but have normal color vision. In contrast, the males who inherit the defective allele will be color blind, because the Y chromosome doesn't have an allele to oppose it.




The term "anti-microbial" is a general term that encompasses drugs, chemicals, or other substances that either kill or slow the growth microbes. These include:
Scottish scientist Alexander Fleming is widely credited with the discovery of the antibiotic properties of penicillin in 1928, although an earlier report had noted the ability of penicillin mold to kill bacteria as early as 1897. There are scattered reports of penicillin mold being used to treat gonococcal infections of the eye of newborns as early as 1930, but it wasn't until the 1940s that penicillin began to be used for clinical infections. It was soon recognized to be a truly remarkable drug. Small doses cured infections caused by Staphylococcus, Streptococcus, Neisseria, syphilis, and many other bacteria. Penicillin use became increasingly widespread and indiscriminate, and resistant strains of bacteria emerged. Increasingly large doses of pencillin were require to cure infections, and some strains were entirely resistant. Other antibiotics were developed, and synthetic penicillin-like drugs were introduced, but strains of bacteria resistant to these newer antibiotics also emerged. At first, these problems were dismissed, but by the 1980s, it had become clear that antibiotic resistance was an important and growing problem. In the 1950s studies were published showing that animals given low doses of antibiotics gained weight more rapidly, and the practice of including antibiotics in grain to promote the growth of cattle, poultry, and swine became widepspread, further compounding the problem of bacterial resistance to antibiotics.
Antibiotics kill or inhibit the growth of susceptible bacteria. Sometimes one of the bacteria survives because it has the ability to neutralize or evade the effect of the antibiotic; that one bacteria can then multiply and replace all the bacteria that were killed off. Exposure to antibiotics therefore provides selective pressure, which makes the surviving bacteria more likely to be resistant. In addition, bacteria that were at one time susceptible to an antibiotic can acquire resistance through mutation of their genetic material or by acquiring pieces of DNA that code for the resistance properties from other bacteria. The DNA that codes for resistance can be grouped in a single easily transferable package. This means that bacteria can become resistant to many antimicrobial agents because of the transfer of one piece of DNA.
Link to more information on antiimicrobial resistance due to antiobiotic use in livestock
Penicillin's ability to kill bacteria was due to it's ability to inhibit a bacterial enzyme that was essential for synthesis of the bacterial cell wall. Penicillin's ability to do this depended on a key structure called a "β-lactam ring". Resistance to penicillin initially occurred as a result of a mutation in a bacterium that created an enzyme (penicillinase) which was capable of breaking down the β-lactam ring. Bacteria that possessed this resistance first evolved in hospitals, but they rapidly spread to the wider community at large. The gene encoding for penicillinase resided not on the bacterial chromosome, but on an extra circlular ring of DNA referred to as a "plasmid." This extra piece of bacerial DNA can be replicated and transferred from a resistant bacterium to one that was previously susceptible by a process referred to as bacterial conjugation.
Methicillin, a chemically modified version of penicillin, was introduced in 1959 to treat infections caused by bacteria resistant to penicillin, but it was effective against a narrower spectrum of bacteria. In addition, strains of Staphylococci resistant to methicillin were reported as early as 1961; these strains had acquired a gene (mecA) wihich inacitivates methicillin by encoding for a protein that binds to it. The mecA gene is carried on an extra "mobile genetic element", the staphylococcal cassette chromosome (SCCmec). Initially, MRSA strains were encountered only in hospitals, but in the late 1990s MRSA was found in the community at large and quickly spread worldwide.
Students in PH709 created a 7-minute public service announcement that explains how MRSA evolved in hospitals and eventually escaped to the community to cause cases of so-called "community-acquired MRSA."
You can learn more about the issues surrounding MRSA and measures to control is by exploring iFrame below which links to the CDC web page on MRSA.
As we noted above, novel genes arise from random mutations, and occasionally such a mutation may confer a bacterium with resistance to an antiobiotic. Once a bacterium has acquired resistance to a particular antibiotic, it passes the resistant allele to subsequent daughter cells that result from binary fission. In addition, bacteria that have acquire a trait such as antibiotic resistance can transfer this allele to other bacteria throught any one of three mechanisms:
When bacterial cells die, they frequently lyse (burst) releasing their intracellular contents, including fragments of DNA, to the environment. These fragments can be taken up and incorporated into the chromosome of a living bacterium to provide the recipient with new characteristics. This process is called bacterial transformation, and if the incorporated DNA contains genes that encode for resistance to an antibiotic, a previously susceptible bacterium can be "transformed" to now be resistant. The video below (23 sec.) provides a quick overview of transformation.
Many bacteria have plasmids, which are small circular pieces of DNA separate from the primary bacterial chromosome. These plasmids can carry genes that provide resistance to antibiotics, and bacteria that contain plasmids are able to conjugate with other bacteria and pass a replicate to recipient bacteria. The electron micrograph below shows two bacteria that are joined by a temporary hollow tube-like connection called a pilus.
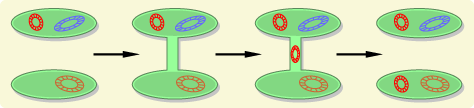
Source: http://evolution.berkeley.edu/evolibrary/article/side_0_0/turboevolution_01
Animation Illustrating Bacterial Conjugation
Source: https://www.youtube.com/watch?v=hm8SZaFmlWg
In 1968 a Shigella epidemic killed 12,500 people in Guatemala. The Shigella bacteria that caused the outbreak had a plasmid carrying resistances to four antibiotics.
Link to CDC web page on Shigella.
Genetic information can also be carried from one bacterium to another by a virus. Bacteriophages (or simply "phages") are small viruses that infect bacteria and use their cellular components to make bacteriophage replicates. During the infection and replication, it is possible for bacterial genes to get incorporated into the viral genome. One of the viral replicates carrying the bacterial allele may then subsequently infect another bacterium and pass the new allele on.
The illustration below begins with a donor bacterium that has a gene (a+) which encodes for resistance to a particular antibiotic. In the 2nd and 3rd scenes the viral proteins and genetic material are replicated and then self-assemble into new viral particles. In the process, however, the bacterial gene for a+ gets incorporated into one of the virus particles. The viral particles are then released from the donor bacterium, and the virus with the a+ gene infects a recipient bacterium, after which the a+ gene can become incoporated tinto the genome of the recipient, which has now acquired this gene for antibiotic resistance.
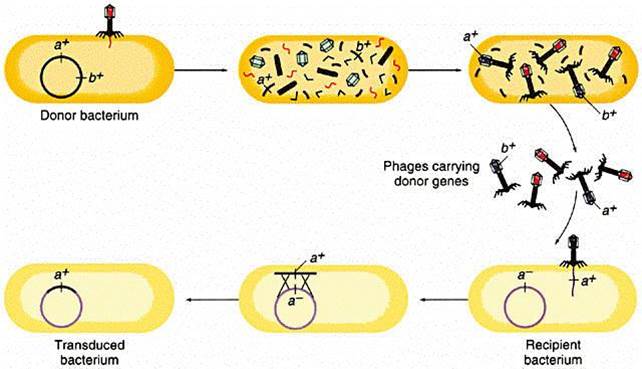
Source: http://www.cbs.dtu.dk/courses/genomics_course/roanoke/genetics980309.html
The development and spread of bacterial resistance to antiobiotics is inevitable, but it could be greatly curtailed through relatively simple measures. These include:
As noted earlier HIV is a retrovirus consisting of a single strand of RNA inside a protein coat. When HIV enters a CD4 lymphocyte, it sheds its protein coat and uses a viral enzyme called reverse trancriptase to create a segment of DNA using the viral RNA as a template. This double-stranded DNA version o HIV then gets incorporated into the DNA of the infected host cell, and this process is called "reverse transcription." This has important consequences for the development of drug resistance by HIV because of several key characteristics of HIV:
Given the persistence of HIV with high rates of replication and high error rates during reverse transcription, mutations in HIV are inevitable, and some of these mutations lead to the eventual development of drug resistance.
HIV infects CD4 lymphocytes and hijacks the cell's machinery to create viral proteins and RNA. HIV's genetic material in RNA, and in order replicate reverse transcriptase results in results in very rapid production of new HIV particles
Two concepts are important to an understanding of the development of drug resistance. First, HIV infection is characterized by high levels of virus production and turnover. In most untreated patients, the total number of productively infected cells in the lymphoid tissue has been estimated to be approximately 107 to 108cells. During the chronic phase of HIV infection, this number is relatively stable, reflecting the balance between the infection of new target cells and their clearance. Because the half-life of infected cells is remarkably short (one to two days), the maintenance of this steady state requires that HIV infect new target cells at a very high rate. Second, the viral population in an infected person is highly heterogeneous. The reverse transcription of viral RNA into DNA is notoriously prone to error,introducing on average one mutation for each viral genome transcribed. Most of these errors are base substitutions, but duplications, insertions, and recombination can also occur. The high rate of HIV infection, combined with the high mutation rate that occurs during each cycle of infection, ensures that patients have a complex and diverse mixture of viral quasispecies, each differing by one or more mutations.
Under these circumstances, it is easy to understand why if any of these mutations can confer some selective advantage to the virus, such as a decrease in its susceptibility to an antiretroviral agent, the corresponding quasispecies will overtake the others, following a simple darwinian selection process.