Genes
Each of our cells has a complete set of our 46 chromosomes, i.e., our entire genome . Altogether our 46 chromosomes contain about 6 billion nucleotides, i.e., 3 billion base pairs. Each chromosome contains thousands of "genes.
. Altogether our 46 chromosomes contain about 6 billion nucleotides, i.e., 3 billion base pairs. Each chromosome contains thousands of "genes. " The segments of DNA that contain genes (referred to as "coding areas") take up only 3-5% of our DNA; the rest of the DNA consists of " non-coding areas ." Altogether our 23 pairs of chromosomes with their 3 billion base pairs carry the code for 20,000-25,000 genes. Most of the genes are transcribed into "messenger RNAs" (mRNA) that provide a template that is used to translate the code into specific proteins. However, about 100 genes are transcribed into "ribosomal RNAs" and "transfer RNAs" that also play a vital role in the synthesis of proteins, which will be described shortly.
" The segments of DNA that contain genes (referred to as "coding areas") take up only 3-5% of our DNA; the rest of the DNA consists of " non-coding areas ." Altogether our 23 pairs of chromosomes with their 3 billion base pairs carry the code for 20,000-25,000 genes. Most of the genes are transcribed into "messenger RNAs" (mRNA) that provide a template that is used to translate the code into specific proteins. However, about 100 genes are transcribed into "ribosomal RNAs" and "transfer RNAs" that also play a vital role in the synthesis of proteins, which will be described shortly.
Transcription and Translation of a Gene
The sequence of bases in DNA can be thought of as the "letters" that provide the basis for the genetic code for all of the proteins synthesized by our bodies, and these, in turn, provide the basis for the structure of all of our cells, all of our enzymes, and all of our inherited traits and characteristics. As noted above, the genetic code is contained in chromosomes which are gigantic molecules of DNA complexed with proteins and wound into a compact structure. Humans have 23 pairs of chromosomes, which carry our entire genome. In eukaryotes chromosomes are located in the cell nucleus, but prokaryotes (bacteria) have a more primitive cellular structure, and they do not have a true nucleus. Instead, the single bacterial chromosome is in the cytoplasm in an area sometimes referred to as the "nucleoid." The production of cellular proteins requires two major processes.
chromosomes are located in the cell nucleus, but prokaryotes (bacteria) have a more primitive cellular structure, and they do not have a true nucleus. Instead, the single bacterial chromosome is in the cytoplasm in an area sometimes referred to as the "nucleoid." The production of cellular proteins requires two major processes.
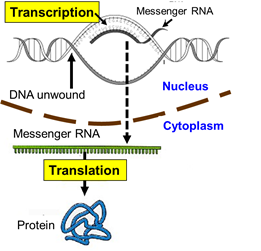
Transcription
First, cellular signals reaching the nucleus cause the TATA-binding protein to the starting point of a particular gene. Additional transcription factors then bind, and an enzyme called RNA polymerase II then binds to the complex. The the polymerase causes the strands of DNA to separate temporarily, and the enzyme synthesizes a strand of messenger RNA (mRNA) to using the the sequence of bases on one strand of DNA (the coding strand) to create a complementary strand of mRNA.
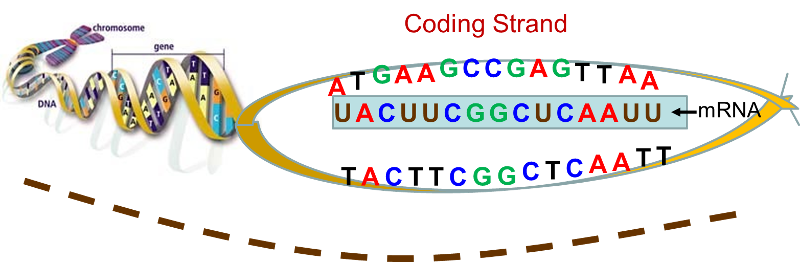
By complementary we mean that the base sequence on mRNA has bases that are the complementary pairs of those on DNA. guanine (G) dictates the insertion of its complement, cytosine (C), and cytosine dictates the insertion of guanine (G); thymine (T) dictates insertion of adenine (A) on mRNA and adenine dictates the insertion of uracil (U). [Note that RNA uses uracil in place of thymine.] Once the strand of mRNA has been created, it leaves the nucleus through pores in the nuclear membrane. The video below gives a fairly detailed picture of the process of transcription. .
Translation
Once the mRNA is in the cytoplasm, it binds to a ribosome, which is composed of protein and a different type of RNA called ribosomal RNA (rRNA). One can think of the ribosome as the work bench where protein is synthesized by covalently bonding amino acids in the sequence specified by the code on the mRNA.
How is the code translated?
One can think of the sequence of bases on mRNA as a series of code letters that are read as a series of three letter "words."
For example, if mRNA had a sequence of bases such as
"...AUGAAGCCGAGUUAAGAU...."
This sequence would, in effect, be read as a series of three-letter words referred to as "codons", each of which specified the insertion of a specific amino acid. In the example just above, the codons or "words" would be:
"...AUG-AAG-CCG-AGU-UAA-GAU...."
Each of these three letter words specifies the insertion of one of the 20 amino acids that make up human proteins. The amino acids are shuttled to the ribosome by a family of transfer RNAs (tRNA), and there are specific tRNAs for each amino acid. The tRNAs consist of a single strand of RNA, but the strand tends to fold back on itself and create loops that are held in place by hydrogen bonds between segments of the tRNA as shown in the illustration below.
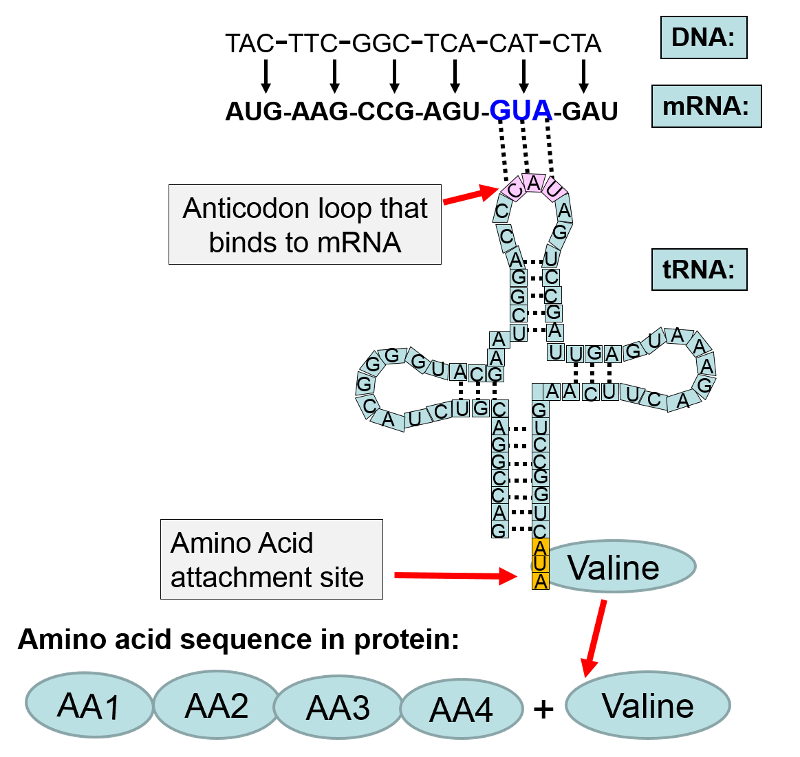
In the illustration above the base sequence CAT on DNA was transcribed to become the codon GUA on messenger RNA. The mRNA left the nucleus and attached to a ribosome where protein synthesis (translation) was initiated. Each codon on mRNA specified a particular amino acid to be added to the growing protein chain. In this example, the first four amino acids are designated as "AA1-AA2-AA3-AA4". The next codon on mRNA was "GUA." The complement to GUA is "CAU" which is the anticodon on a transfer RNA that carries the amino acid valine. The anticodon CAU on the tRNA for valine bonded to the GUA codon on mRNA. This positioned valine as the next amino acid in sequence, and with the addition of cellular energy (ATP), valine became covalently bonded to AA4 in the amino acid chain.
In the section above on transcription, we focused on creating the mRNA for a specific gene; those events took place in the cell nucleus. The figure below illustrates the subsequent events that take place after mRNA leaves the nucleus and attaches to a ribosome and initiates translation.

Sixty-one codons specify an amino acid, and the remaining three act as stop signals for protein synthesis. For example, in the figure below the codon UGA signals an end to synthesis of the protein. The code for all possible three-letter codons on mRNA is shown in the blue table below. Note that there is some redundancy in the code. For example, there are four separate codons for the amino acid proline. Nevertheless, the code is unambiguous, because no triplet codes for more than one amino acid. In addition, with only a few minor exceptions, the same code is universally found in viruses, bacteria, protists, plants, fungi, and animals.
Note that in the example below, UGA, is a signal to STOP, meaning that the amino acid chain is complete and no more amino acids are to be added. Bear in mind that these illustrations include just short sequences of codons, and an actual protein would generally have a much longer sequence. Nevertheless, these examples illustrated how the code is transcribed from DNA to mRNA and how the mRNA is then translated in order to specify the sequence of amino acids in a particular protein which is the product of that particular gene on a chromosome.
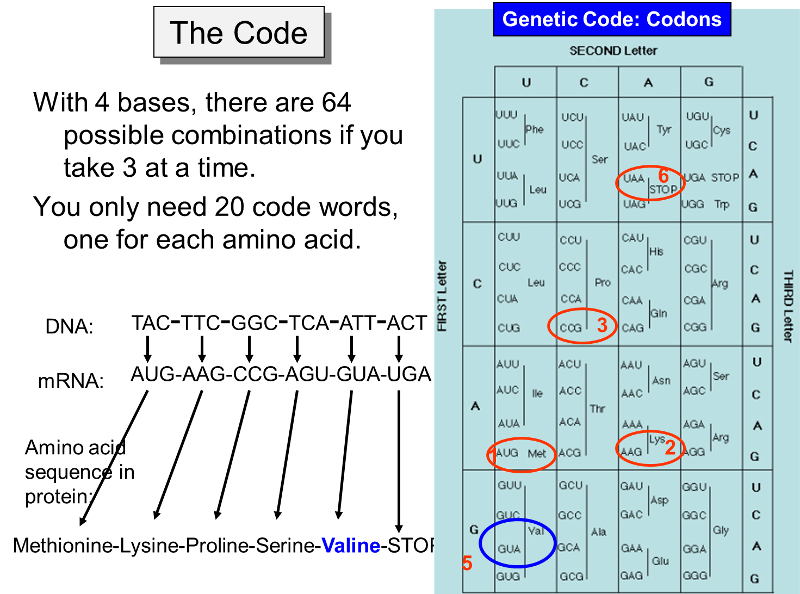
The figure above makes it clear that the order of the codons within a gene (a segment of DNA encoding for a specific protein) specifies the amino acid sequence in the protein. The start signal for protein synthesis is the codon AUG, which specifies incorporation of the amino acid methionine. When the mRNA attaches to a ribosome, enzymes look for the AUG codon, not only as a start signal, but also as a means of knowing exactly which is the first letter of each of the three-letter codons. For example, a messenger RNA might have the sequence of codons shown in the illustration above, i.e.,
AUG-AAG-CCG-AGU-GUA-UGA-... etc.
However, if the signal to start was shifted by one nucleotide (e.g., starting at the first "U" instead of the "A"), the codons would be read as:
UGA-AGC-CGA-GUG-UAU- ..etc.
and this would result in synthesis of an very different sequence of amino acids. Errors in the sequence of amino acids can, in fact, result from mutations, as described below.
The video below gives a detailed summary of the events that occur during translation of the mRNA template to a protein.
This next video is an excellent illustration of transcription and translation, but it illustrates these in a way that provides a real-time approximation.
|
An Interesting Variation by HIV
The human immunodeficiency virus (HIV) is known as a retrovirus. It consists of a single strand (molecule) of RNA inside a protein coat. When HIV binds to a T lymphocyte it enters the lymphocyte and sheds its protein coat. A viral enzyme called reverse trancriptase then uses the strand of viral RNA as a template to create a molecule of DNA which can become incorporated into the DNA of the infected host cell. In this case, RNA is being used to create a molecule of DNA, and the process has been dubbed "reverse transcription."
|
Mitochondrial DNA
Most of the DNA in eukaryotic cells is contained in the chromosomes within the membrane-bound nucleus, but the mitochondria also have small amounts of DNA (mitochrondrial DNA or mtDNA). As you will recall, mitochondria are membranous subcellular organelles within which there are chains of enzymes that generate cellular energy in the form of ATP (adenosine triphosphate) through a process called oxidative phosphorylation. In addition to their role in production of ATP, mitochondria also regulate apoptosis (programmed cell death) [for more on apoptosis see the section on Apoptosis in the module on the Biology of Cancer]. Mitochondria also play a role in the synthesis of cholesterol and heme (a component of hemoglobin, the oxygen carrying molecules in red blood cells). Mitochondrial DNA consists of 37 genes. Thirteen of these provide the genetic code for synthesizing the enzymes involved in oxidative phosphorylation, and the rest encode the transfer RNAs (tRNA) and ribosomal RNA (rRNA) required for synthesis of the enzymes for oxidative phosphorylation.
Inherited mutations in mitochondrial DNA can also cause a variety of problems with growth, development, and function throughput the body as a result of impaired ability to generate ATP. These conditions can produce muscle weakness and wasting, diabetes, kidney failure, heart disease, dementia, hearing loss, visual problems. In addition, mitochondria can also undergo somatic mutations (non-inherited) which may contribute to aging and age-related diseases.
Additional Resources for Mitochondrial DNA
- Molecular Expressions, a web site from the Florida State University Research Foundation. Link to their illustrated introduction to mitochondria and mitochondrial
DNA . - Link to an overview of mitochondrial
DNA from the Neuromuscular Disease Center at Washington University. - The Howard Hughes Research Institute: Link to an article about recent research into mitochondrial
function .
For more information about conditions caused by mitochondrial DNA mutations:
- Genetics Home Reference provides background information about mitochondria and mitochondrial DNA written in consumer-friendly language.
- The Cleveland Clinic's Introduction to mitochondrial
disease . - An overview of mitochondrial
disorders from GeneReviews.