
Basic Cell Biology
The short video segment below (3:36) is from Discovery Channel. It's a very basic introduction to cells, but may be of interest to students with little background in the sciences.
Source: http://youtu.be/u54bRpbSOgs
After successfully completing this section, the student will be able to:

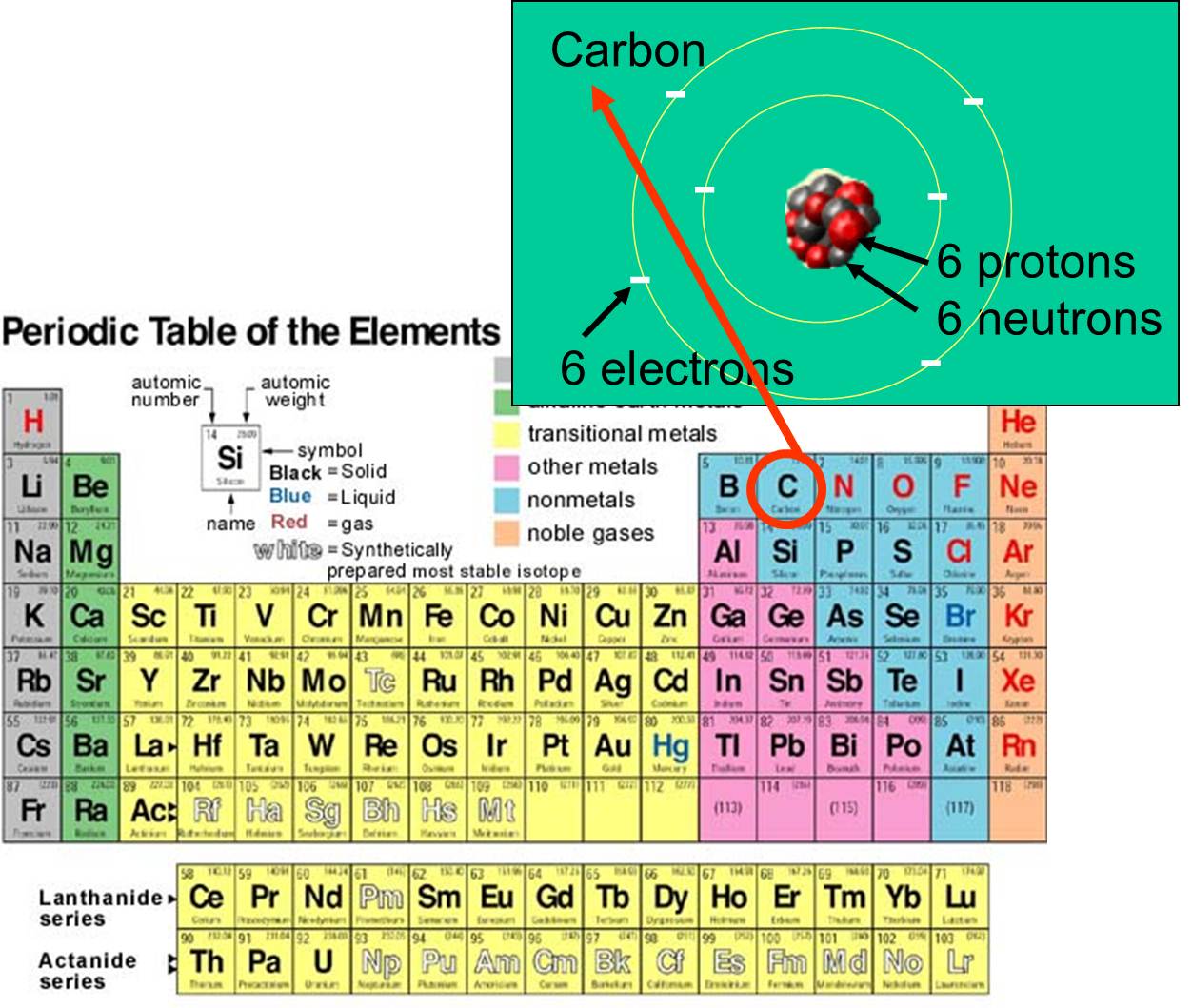 All matter, whether it is living or not, is composed of chemical elements; these are fundamental chemicals in the sense that they are what they are - they can't be changed into another element. Each element is distinguished by the number of protons, neutrons, and electrons that it possess. For example, carbon's atomic number is 6, and has an atomic mass of about 12, because it has 6 positively charged protons and 6 non-charged neutrons. The 6 charged electrons contribute very little to the atomic mass. There are 92 naturally-occurring elements on earth. The array of elements and their subatomic structure are summarized by the periodic table of the elements, shown to the right.
All matter, whether it is living or not, is composed of chemical elements; these are fundamental chemicals in the sense that they are what they are - they can't be changed into another element. Each element is distinguished by the number of protons, neutrons, and electrons that it possess. For example, carbon's atomic number is 6, and has an atomic mass of about 12, because it has 6 positively charged protons and 6 non-charged neutrons. The 6 charged electrons contribute very little to the atomic mass. There are 92 naturally-occurring elements on earth. The array of elements and their subatomic structure are summarized by the periodic table of the elements, shown to the right.
In living organisms the most abundant elements are carbon, hydrogen, and oxygen. These three elements along with nitrogen, phosphorus, and a handful of other elements account for the vast majority of living matter. An atom is one single unit of a chemical element. Some of these elements that are abundant in organic molecules are shown below.
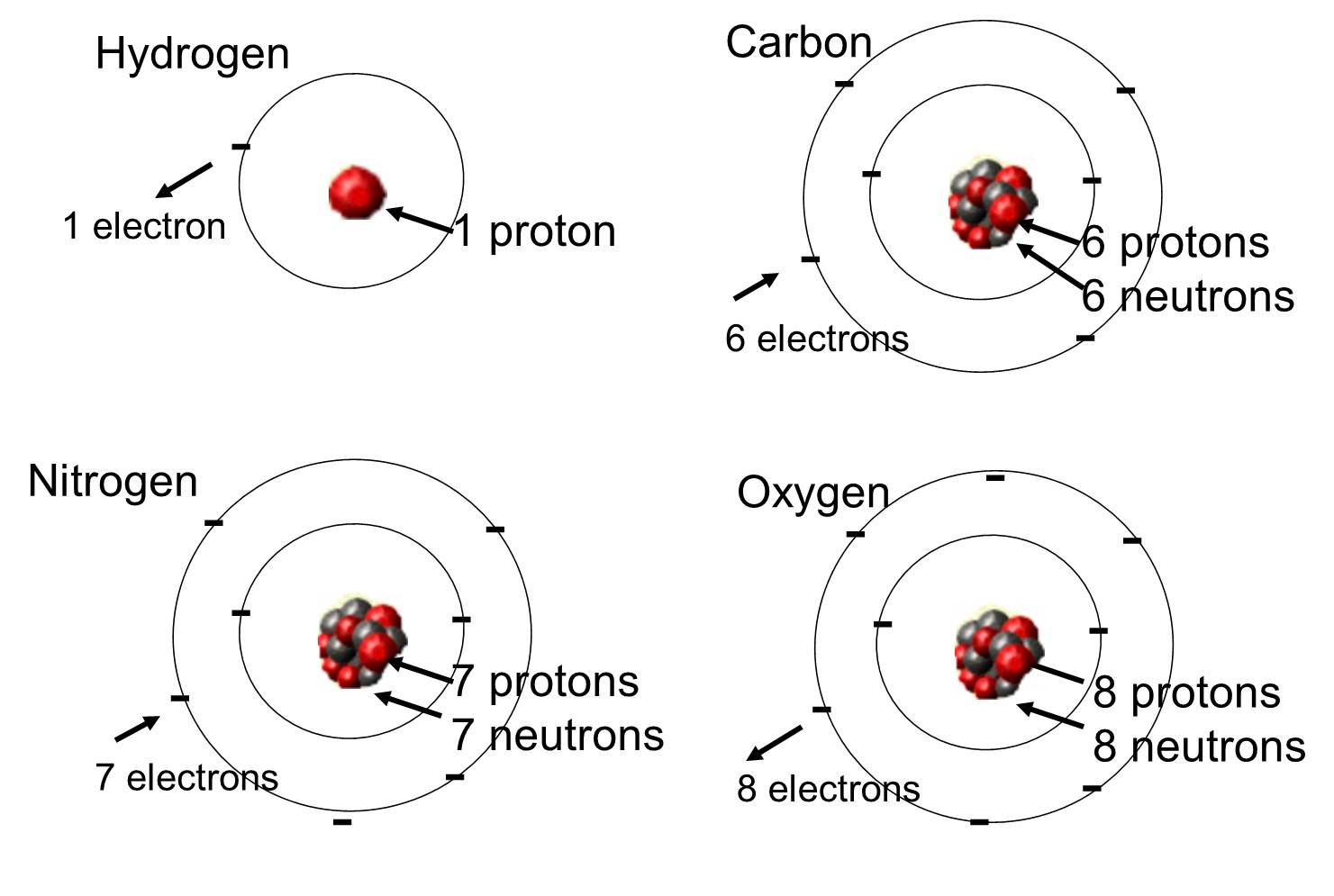
 Atoms can combine with other atoms by forming chemical bonds.
Atoms can combine with other atoms by forming chemical bonds.
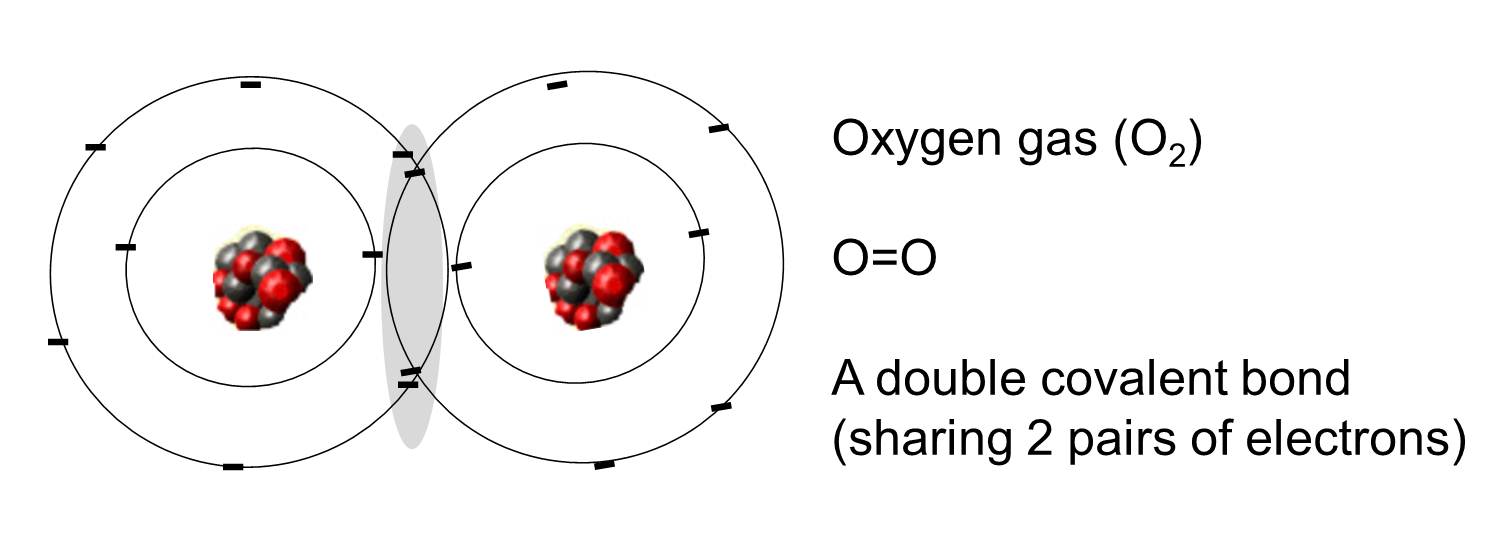
A covalent bond is one in which one or more pairs of electrons are shared by two atoms. The illustration to the right shows two atoms of oxygen that are covalently bonded by the sharing of two pairs of electrons as illustrated in the shaded area.
The figure below shows a series of molecules formed by covalent binding. Mouse over each molecule to see a brief description.
,
Note also that the sharing of electrons is not always equal. For example, in a water molecule, the negatively charged electrons spend more time in the vicinity of the heavier oxygen atom.
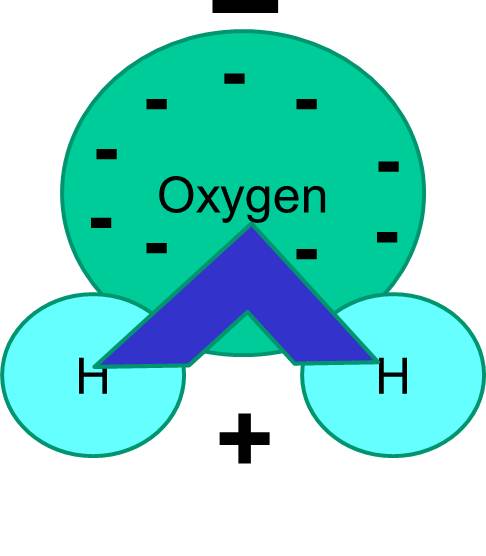
The net result is that the water molecule has one end that is more negative relative to the other end. Water is therefore a "polar" molecule. We will see that this polarity has important implications for many biological phenomena including cell structure. You may have heard the expression "like dissolves like." What this means is that polar molecules dissolve well in polar fluids like water. Sugars (e.g., glucose) and salts are polar molecules, and they dissolve in water, because the positive and negative parts of the two types of molecules can distribute themselves comfortably among one another.
Sodium has a single electron in its outermost orbital shell, and it is thermodynamically more stable if it gives up this electron. This loss of a negative electron results in a positively charged sodium ion, abbreviated Na+. Chlorine, on the other hand, has seven electrons in its outermost orbital shell, and it is more thermodynamically stable if it acquires an extra electron to complete the outer orbital shell. This results in a negatively charged chloride ion, abbreviated Na+. The positively charged sodium ions and the negatively charged chloride ions attract each other and result in the formation of an ionic bond. In the absence of water, sodium and chloride form a crystal lattice because of the attraction of negative and positive ions.
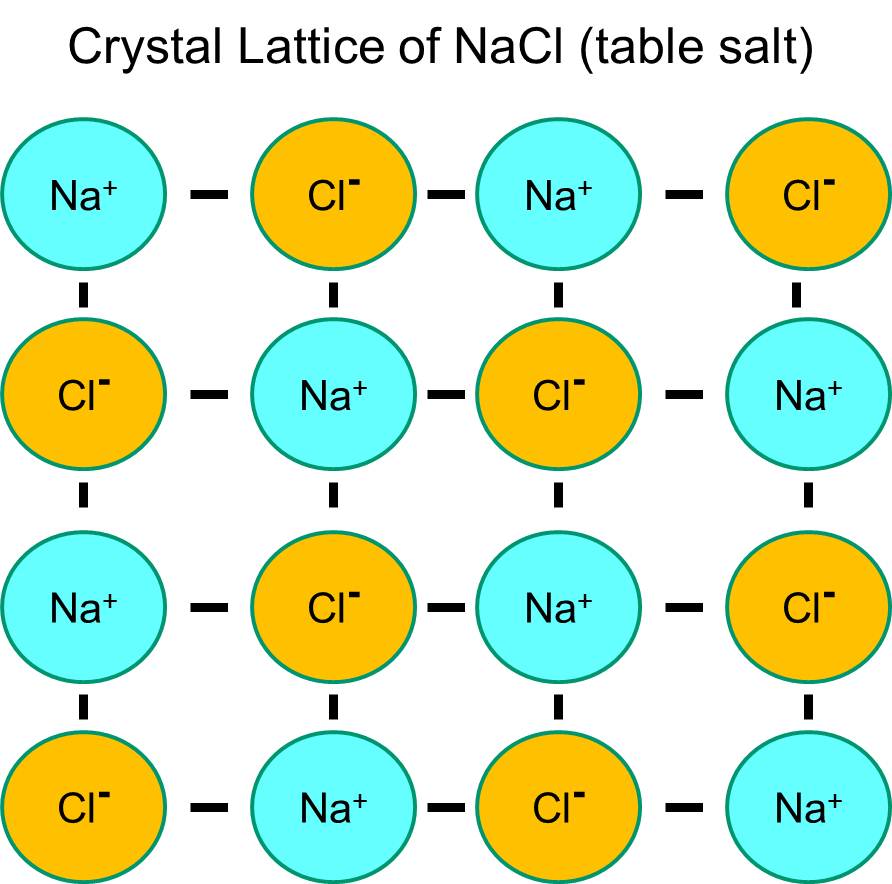
However, if sodium chloride crystals are placed in water, the polar water molecules will "hydrate" the sodium and chloride atoms because the water molecules are polar. In the illustration below the darker blue V-shaped figures represent water molecules, which are polar. The positive ends of the water molecules are attracted to the negatively charged chloride ions, while the negative pole of the water molecule is attracted to the positive sodium ions. As a result, the ions are hydrated and the crystal lattice dissolves into the aqueous solution. This is exactly what happens when you add crystalline table salt to a glass of water.
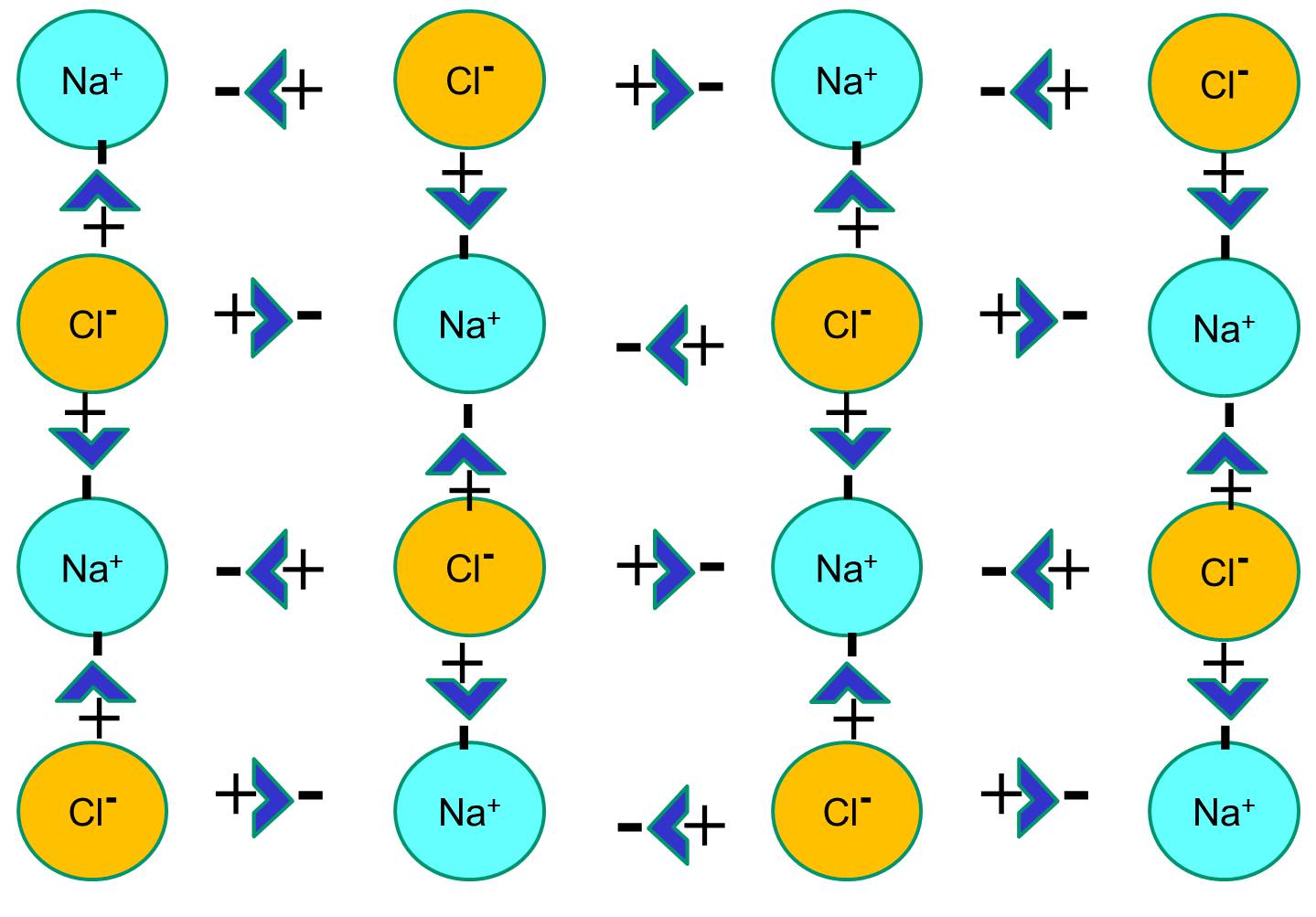
The video below provides an animated explanation of how salts like NaCl dissolve in water.
More Complex Biological Molecules
The stuff of life is amazingly diverse and complex, but it is all based on combinations of simple biological molecules. Biological molecules are often made from chains & rings of carbon. These molecular structures can be represented by "stick drawings" that show the component atoms (e.g., C, H, N, O for carbon, hydrogen, nitrogen, and oxygen respectively) and show the bonds between them as dashes. A single dash ( - ) represents a single bond, and a double dash (=) represents a double bond.
Note that some common "groups" are depicted without showing the bonds between them. For example, the hydroxyl group (-OH) consists of a hydrogen atom bonded to an oxygen atom:
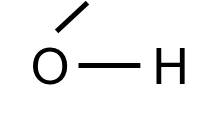
The hydoxyl group will commonly be bonded to a carbon atom in this fashion:
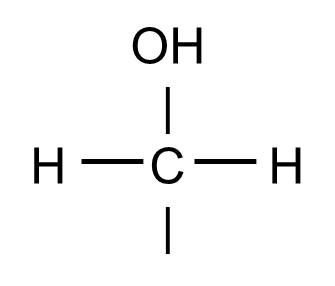
And this structure might be found, for example, as part of a glucose molecule, depicted below.
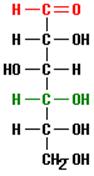
This molecule of the sugar glucose consists of 6 carbon atoms bonded together as a chain with additional atoms of oxygen and hydrogen. Note that the previous structure (a carbon to which two hydrogens and one hydroxyl group are bound) is located at the bottom of this glucose chain where it is written using the notation CH2OH.
This glucose chain forms a ring in aqueous solutions, e.g., in body fluids, as shown below.
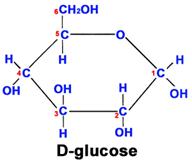
Fructose is another sugar that also has 6 carbons, 12 hydrogens, and 6 oxygen atoms. However, the arrangement of the atoms is different, and this makes it much sweeter than glucose and also affects its ability to combine with other molecules.
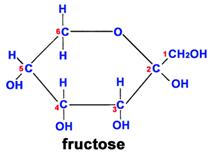
Another important theme is that single units of biological molecules (monomers) can join to form increasingly complex molecules (polymers). For example, two monosaccharide sugars can also become bound together chemically to form a disaccharide. Sucrose is the disaccharide in common sugar that we buy at the grocery store. The structure of sucrose is shown below.
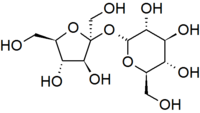
Glucose and fructose are examples of monosaccharides, meaning they consist of a single sugar unit, while sucrose is an example of a disaccharide. However, sugar units can be bonded or linked together to form polysaccharides, which consist of many sugars linked together to form extensive chains of sugars. Plants store energy as starch, which consists of very long chains of glucose linked together.
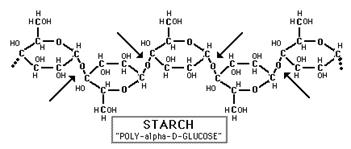
Source: http://www.science-projects.com/Amylase.htm
Animals store energy as glycogen, which consists of more highly branched chains of glucose. Collectively, sugars, starch, and glycogen are know as carbohydrates, and they are an important source of cellular energy.
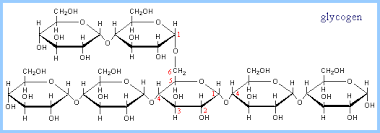
Source: https://www.rpi.edu/dept/bcbp/molbiochem/MBWeb/mb1/part2/glycogen.htm
Cellulose is yet another polysaccharide formed from glucose. Cellulose is composed of unbranched, parallel chains of glucose. A key feature is that the chains bond to one another to form strong fibers that serve a structural purpose. Humans do not have the enzymes necessary to break the bonds in cellulose, and any cellulose we ingest passes through our digestive systems. It is a major component of what we refer to as dietary "fiber."
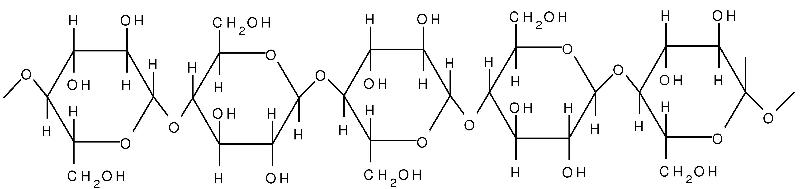
Source: http://www.intechopen.com/books/cellulose-fundamental-aspects/cellulose-microfibril-angle-in-wood-and-its-dynamic-mechanical-significance
Lipids are a family of compounds whose diversity is also made possible by building complex molecules from multiple units of simpler molecules, and once again one sees characteristic rings and chains.
You have probably heard the expression "oil and water don't mix," and you have observed how salad dressing composed of vinegar (which is aqueous, i.e., largely water) and oil will separate when left to stand. This incompatibility is due to the fact that water molecules are polar, but oil is non-polar. Water is a polar molecule because the negatively charged electrons that spin around the nuclei of the atoms are not evenly distributed. The oxygen atom has much more mass than the two hydrogen atoms, and therefore the electrons spend more time in the vicinity of the oxygen atom. As a result, the end of the water molecule where oxygen is located is relatively negative in charge, whereas the end with hydrogens is relatively positively charged. The positive ends of the water molecule are attracted to the negative ends of adjacent water molecules, as shown in the figure below, and this enables water molecules to coalesce. You may have also seen water bead on a car windshield as a result of this phenomenon.
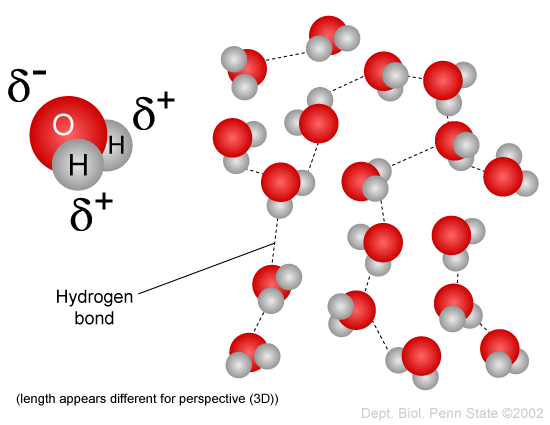
Source: http://www.personal.psu.edu/staff/m/b/mbt102/bisci4online/chemistry/chemistry3.htm
Lipids, i.e., fatty molecules, on the other hand, are non-polar, meaning that the charge distribution is evenly distributed, and the molecules do not have positive and negatively charged ends.. Non-polar molecules do not dissolve well in polar solutions like water; in fact, polar and non-polar molecules tend to repel each other in the same way that oil and water don't mix and will separate from each other even if they are shaken vigorously in an attempt to mix them. This distinction between polar and non-polar molecules has important consequences for living things, which are composed of both polar molecules and non-polar molecules. The next sections will illustrate the importance of this.
Fatty acids are chain-like molecules that are important components of several types of lipids. The illustrations below show two different fatty acid molecules. Each has a characteristic carboxyl group (the -COOH) attached to a chain of carbons with hydrogen atoms attached to the carbon chain. Two things are noteworthy. First, the hydrocarbon chain is very non-polar and therefore doesn't dissolve in water very well. However, hydrocarbon chains do associate with each other readily. Second, note that the unsaturated fatty acid has two hydrogens removed, and this allows formation of a double bond, i.e., a stronger bond between two of the carbon atoms. Note also that the double bond tends to produce a bend or a kink in the fatty acid. The illustration to the right shows two other common fatty acids: stearic acid, which is a straight 18 carbon chain with no double bonds, and oleic acid, which is an 18 carbon chain with a single double bond, which cause a bend in the carbon chain.
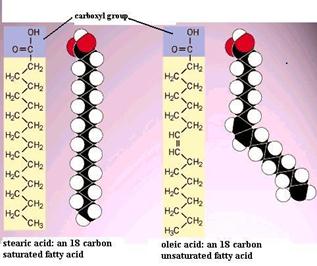
A fat molecule is a type of lipid that consists of three fatty acid molecules connected to a 3 carbon glycerol backbone, as shown on the right. The three fatty acids can be different from one another. Since the hydrocarbon chains are very non-polar, fats do no dissolve in water; instead, fat molecules tend to coalesce with one another. Since a fat molecule has 3 fatty acids connected to a glycerol molecule, they are also called trigylcerides.
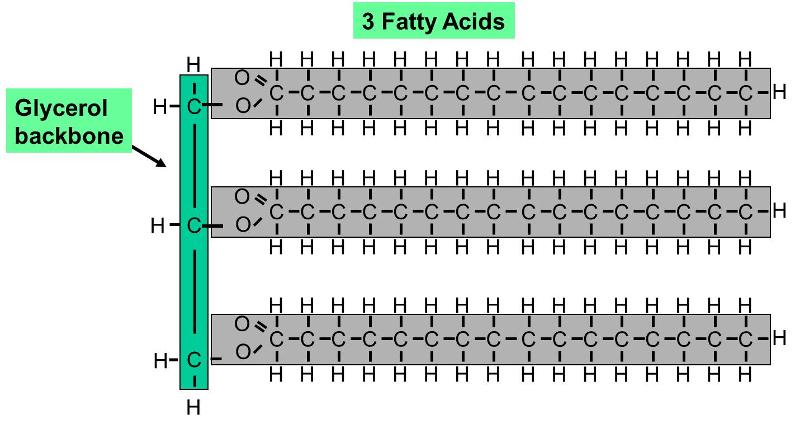
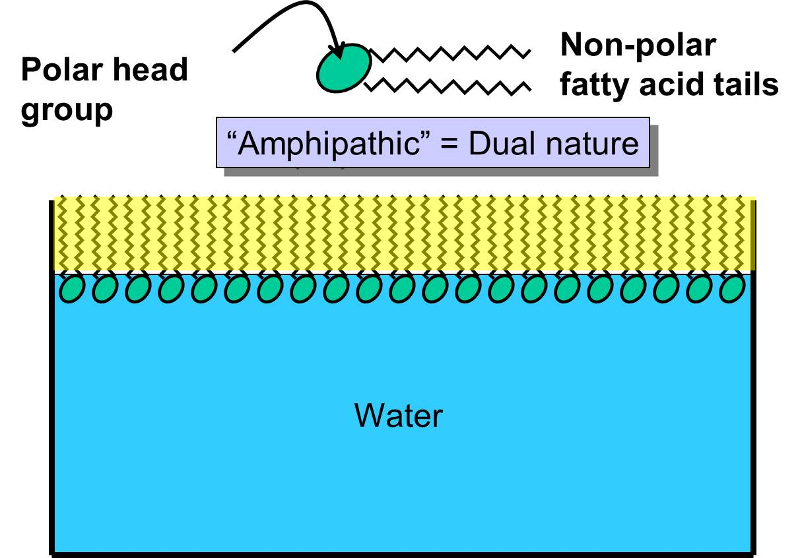
Phospholipids constitute another important class of lipids. These are similar to similar to trigylcerides in that they have a glycerol backbone, but there are only two fatty acids connected to glycerol. The third carbon of the glycerol backbone is attached to a phosphate group (an atom of phosphorus bonded to four atoms of oxygen), and the phosphate group is attached to a base molecule of choline, serine, or ethanolamine. The part of the phospholipid with phosphate and the base is actually very polar, and it tends to rotate away from the two fatty acids. This makes phospholipid molecules have a hairpin shape. The head of the hairpin is very polar and therefore likes to associate with water (it is hydrophilic), while the two fatty acid chains (the "tails") are very non-polar and tend to avoid water (hydrophobic) and associate with other hydrocarbon chains.
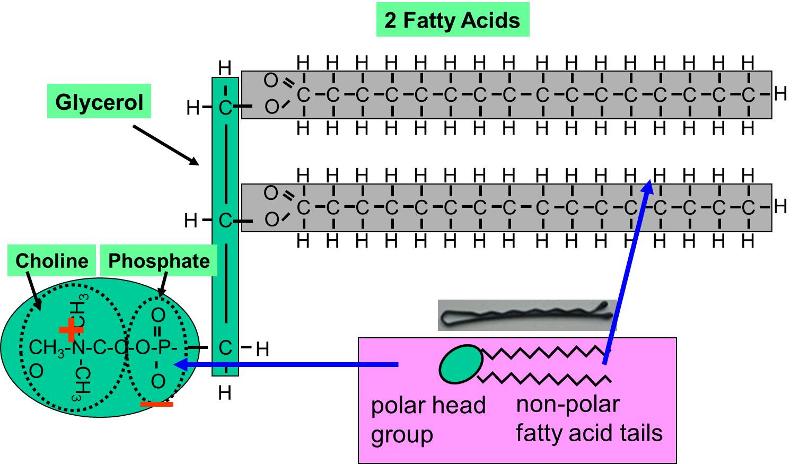
Phospholipids can be described as amphipathic ("amphi" means "both"), because they have this dual nature (part polar and part non-polar). This characteristic causes phospholipids to self-associate into large macromolecular complexes in an aqueous (watery) environment.
Cholesterol is also an important component of animal membranes (plant membranes have a similar, but distinct 'sterol' in their membranes). It is a lipid, because it is composed almost entirely of carbon and hydrogen, but it is different from fatty acids, fats and phospholipids in that it is arranged in a series of rings. The rings consist of 5 or 6 carbon atoms bonded together. The carbon atoms at the apices of the hexagonal and pentagonal rings have hydrogen atoms attached to them. The ring-like structures are fairly rigid, but there is also a hydrocarbon tail, which is somewhat flexible. The entire structure is somewhat reminiscent of a fancy kite with a tail.
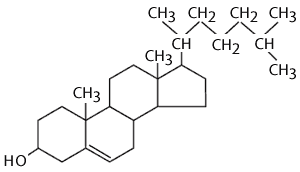
Cholesterol is very non-polar, except for the hydroxyl group attached to the first ring. Consequently, in an animal cell membrane the polar hydroxyl group sticks into the aqueous environment (either extracellular water or intracellular water), and the rest of the cholesterol molecule, which is non-polar, is found among the non-polar fatty acid tails of the phospholipids.The image below depicts a section of a cell membrane with water outside and inside. The polar head groups of the phospholipids are represented in red, and their non-polar fatty acid tails are shown as zig-zag lines extending from the polar head group. As we we see in greater detail, cell membranes consist of a bilayer of phospholipids with other molecules inserted into the bilayer. This illustration shows five cholesterol molecules (the black structures with four conjoined rings) inserted into the lipid bilayer. Most of the cholesterol molecule in non-polar and therefore associations with the non-polar fatty acid tails of the phospholipids. However, the hydroxyl group (-OH) on cholesterol carries a negative charge and therefore associates with the polar environment of water either inside the cell or outside.
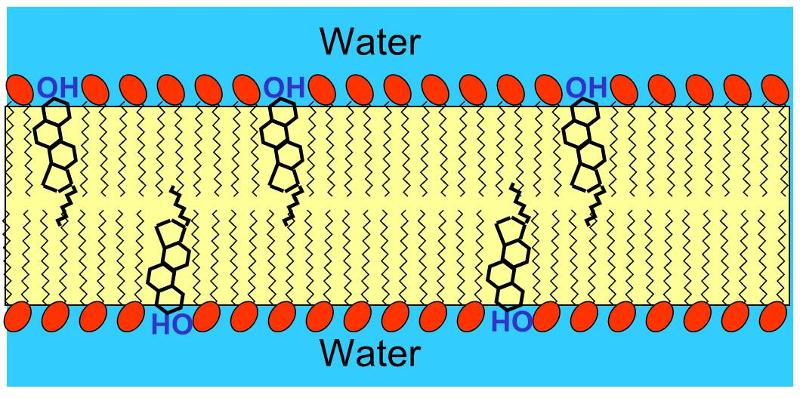
If you were to add small amounts of phospholipid molecules to water, they would float to the surface and align so that the polar head groups awere in the water, and the non-polar fatty acid tails would stick up from the surface of the water and form an oily film. If you were to continue to add phospholipids, the film would eventually cover the entire surface.
Once the surface of the water becomes completely saturated with phospholipid, addition of still more phospholipid would result in formation of a bilayer within the water (as shown on the left), since this would be the most thermodynamically stable structure, allowing all of the polar heads of the phospholipids to be in contact with water, while at the same time allowing all of the non-polar fatty acid tails to be sheltered amongst themselves in an oily layer that is away from the water.
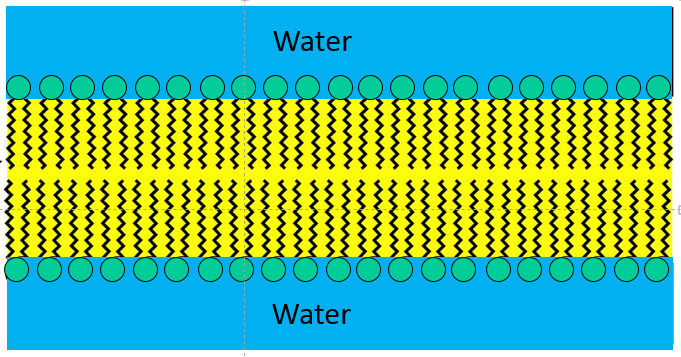
With the addition of even more phospholipid to this aqueous solution the phospholipids would spontaneously form spherical bilayers of phospholipids that had water inside and outside as depicted in the figure below.
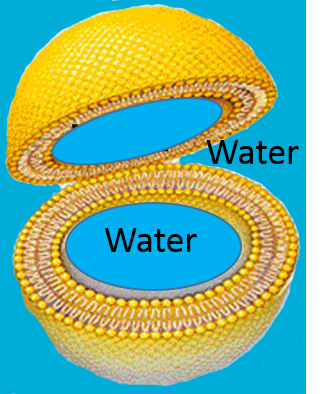

This bilayer structure is actually the basic structure for cell membranes and many of the internal structures (organelles) within cells. Imagine a cell as a three dimensional sac consisting of a bilayer of phospholipid molecules. There is water inside the cell and outside the cell, and the polar heads of the phospholipids protrude into the water (shown in blue). Certainly the structure of cells is far more complex than this. Cell membranes have many proteins and glycoproteins which serve many functions,, e.g. as signal receptors and transport conduits to move molecules in and out of the cell. The video below gives a sense of the structure and function of the plasma membrane, or cell membrane, and depicts it as a "fluid mosaic."
Source: http://www.youtube.com/watch?v=owEgqrq51zY
The Flash animation below gives further detail about the functions of some of the membrane's proteins. Click on the name of each protein type to see more detailed information. [This Flash animation is from "Biology - The Unity and Diversity of Life", 9th edition, by Cecie Starr and Ralph Taggart, Brooks/Cole - Thomson Learning, 2001.]
There are two types of nucleic acids that are important to living things.
These molecules are also polymers of smaller units called nucleotides; each nucleotide consist of a sugar (ribose or deoxyribose), a phosphate group, and one of several "bases" that are either purines or pyrimidines. Alternating sugar molecules and phosphate groups are bonded together to form the backbone of the nucleic acid, and a purine or pyrimidine base is bonded to each of the sugars, as illustrated below.
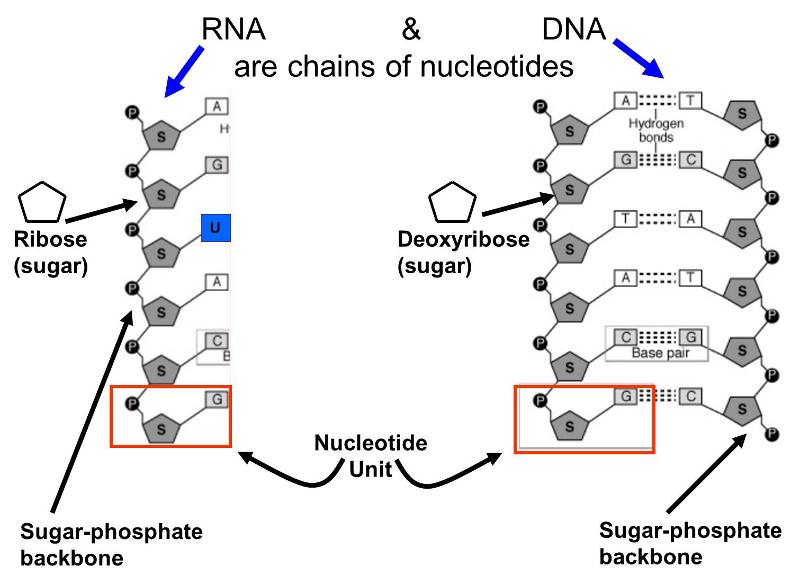
There are several differences between DNA and RNA.
The cells of living organisms have chromosomes which contain an inherited code for synthesizing all of the proteins that the organism produces. In essence, each chromosome is a gigantic molecule of double stranded DNA wound tightly into a double helix. A single chromosome contains thousands of genes, segments of DNA that encode for specific proteins. In a highly regulated process, cellular enzymes can unwind a particular segment (gene), and other enzymes move along a gene using one strand of DNA as a template to synthesize a complementary strand of messenger RNA. This newly synthesized messenger RNA will then leave the cell nucleus and move to the cytoplasm of the cell where the RNA will in turn be used as a template to synthesize a specific protein.
This process will be clearer when we explore it in more detail in another online module. For the time being the video below provides an overview of this process that will be helpful.
This short animation from the Discovery Channel provides a nice overview of the transcription and translation. Transcription is the process by which a gene, segment of DNA that encodes for a specific protein, serves as a template for the synthesis of a messenger RNA (mRNA) for that specific protein. Transcription takes place inside the cell nucleus where chromosomal DNA is located. The mRNA then leaves the nucleus through special pores in the membrane of the nucleus. Once the mRNA emerges from the nucleus, it attaches to a two part structure called a ribosome, which consists of ribosomal RNA (rRNA). Enzymes also attach to the ribosomal complex and aid in the process of translation, in which the coded sequence of bases on the mRNA is translated and directs the synthesis of a chain of amino acids, which are the building blocks of proteins.
Source: http://youtu.be/1fiJupfbSpg
Proteins are another class of enormously diverse organic molecules that are made from multiple units of simpler molecules arranged in chains. All proteins are made from combinations of the 20 amino acids show below. As shown below, each of these 20 amino acids has a central carbon (the alpha carbon) bonded to an amino group (-NH2 i.e., nitrogen bonded to two hydrogens) at one end and a carboxyl group ( -COOH) at the other end.
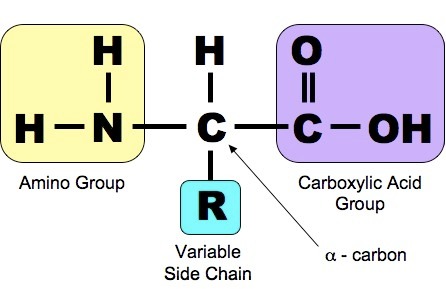
Source: http://study.com/academy/lesson/threonine-amino-acid-structure-function.html
What distinguishes one amino acid from another is the side chain of atoms that is also bonded to the alpha carbon (designated "R-group on the right).
.gif)
Source: http://quotesgram.com/amino-quotes/
The primary structure of proteins results from linking together various combinations of these 20 amino acids with peptide bonds, which link the carboxyl group of one amino acid to the amino group of another amino acid.
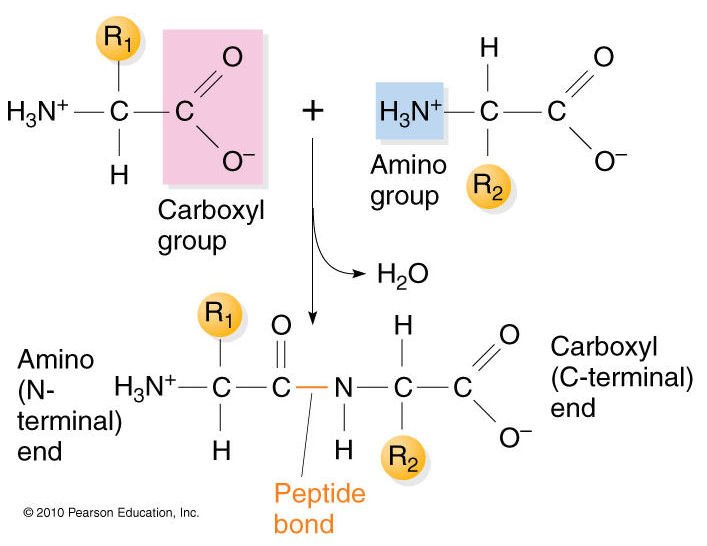
Source: https://biochemistry3rst.wordpress.com/tag/proteins/
Now imagine that dozens or even hundreds of amino acides are linked together in chains of varying length to create the primary structure of a protein. Proteins are sometimes referred to as polypeptides because they consist of chains of amino acids linked together with peptide bonds.
The chains of amino acids establish the primary structure of a protein, but interactions (both attractive forces and repulsive forces) among the components of the chain reshape the protein into its filnal three-dimensional structure. These attractive and repulsive forces among the amino acids in the chain cause segments of the chain to take on one of several characteristic forms that provide the secondary structure of the protein. For example, the hydrogen on the amino group of one amino acid can form a weak "hydrogen bond" to the oxygen atom in the carboxyl group of another amino acid elsewhere on the chain. Hydrogen bonding can cause portions of the polypeptide chain to form zig-zag sections called "beta sheets" (which are very prominent in the protein fiber in silk, for example), and it can also cause sections of the polypeptide to twist into a cork screw-shaped structure called an "alpha helix." Other sections of a polypeptide may be referred to as "random coils" because they fold but do not have a regular structural shape.
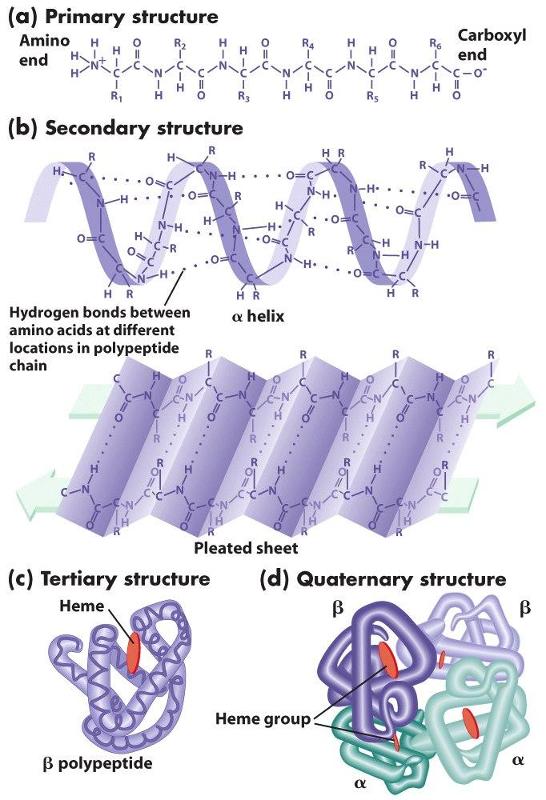
Proteins also have a tertiary level of structure as a result of ionic, hydrogen, or covalent bonds between the "-R" groups of the amino acids. As a result, alpha helical segments, beta pleated sheets, and random coils fold upon themselves. Folding and placement in a cell will also be influenced by the polarity of the amino acids. Some amino acids have side chains that are polar and others have non-polar side chains. If some sections of the chain contain mostly non-polar amino acids, while other sections contain mostly polar amino acids, the non-polar sections will self-associate in the interior of the molecule away from water, and the polar sections will be arrayed on the exterior of the molecule.
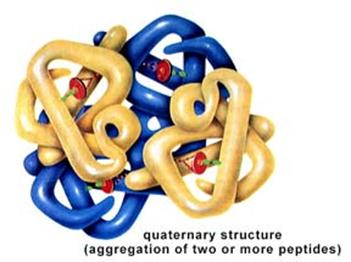
Finally, quaternary structure refers to the association of two or more polypeptide chains or subunits into a larger entity. For example, the hemoglobin molecule (shown in (d) to the left) consists of two alpha subunits and two beta subunits; each of these four polypeptide chains has a binding site for oxygen. Transport proteins in cell membranes frequently consist of multiple subunits as well.
The three dimensional structure of proteins goes hand in hand with their function. Moreover, the three dimensional shape of a protein (its conformation) may change depending on changes in its local environment, and this also may relate to its function. To illustrate, consider the function of an enzyme whose purpose is to cleave the phosphate groups from a molecule called cyclic AMP. The enzyme is depicted in the figure to the right. The drawings on the left side of the figure show the enzyme folding into its quaternary conformation (folded protein), and the drawing on the right is a close up of the binding site, showing a molecule of cyclic AMP (pink shading) nestled in the arms of the binding site. Chemical groups on the cyclic AMP (the substrate) are interacting with chemical groups on the enzyme through ionic and hydrogen bonds. The binding site is specific for cyclic AMP, which fits into the protein binding site in much the same way that a key fits into a specific lock. This interaction then causes the conformation of the enzyme to change, and this bends the cyclic AMP in a way that facilitates cleavage of the phosphate group. Once this occurs, the two resulting products are released, and the enzyme reverts back to its resting conformation.
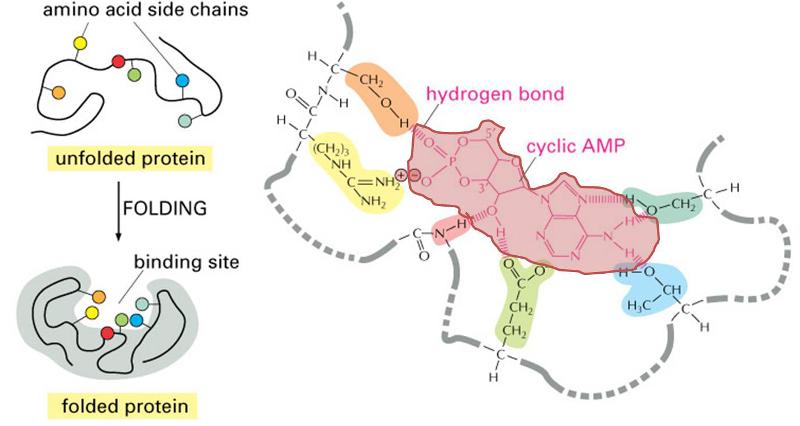
(Illustration adapted from http://accessexcellence.org/RC/VL/GG/ecb/protein_binding_site.php )
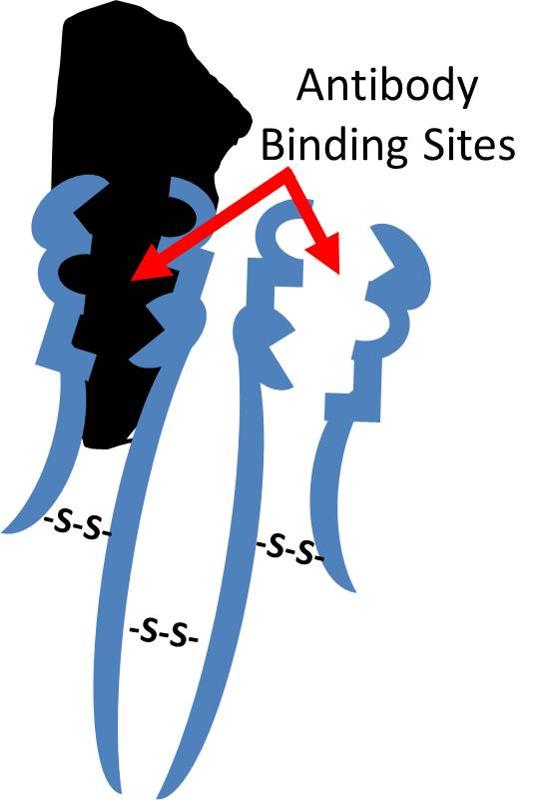
The three dimensional shape of proteins and this concept of a specific binding site is relevant not only for the interaction of enzymes and their substrates, but also for receptors which bind chemical signals in a specific way, e.g,, a protein receptor embeded in the cell membrane that has a complementary shape to the signal molecule histamine receptor as illustrated below. Binding of histamine molecules to their corresponding receptors causes a change in the conformation of the proein receptor that triggers a series of biochemical responses within the cell, such as contraction of smooth muscle cells in thr bronchi of a child having an asthma attack.
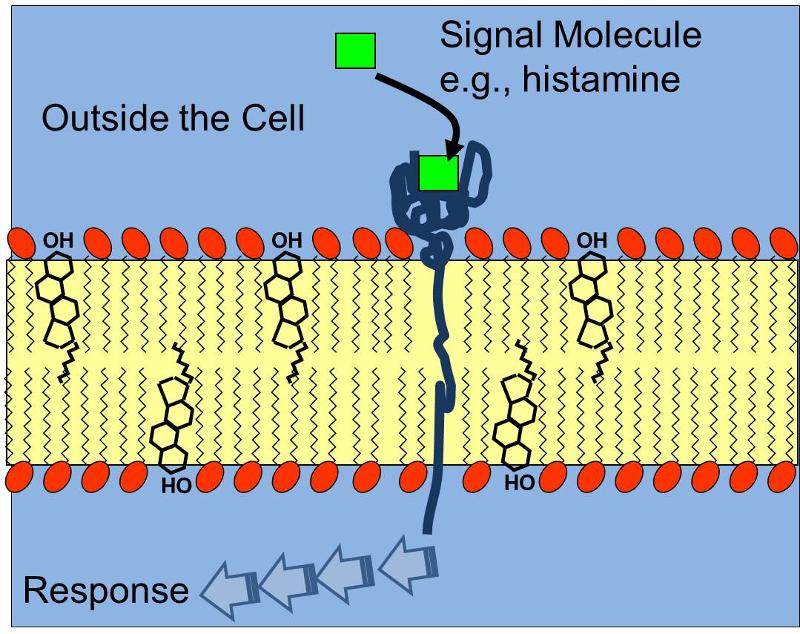
Antibodies that are produced in response to infections have binding sites that enable them to bind to complementary shapes on foreign proteins. The illustration below shows an IgG antibody in blue, composed of two short polypeptide chains and two long chains that are held together with disulfide bonds (S-S).The two identical binding sites on this antibody have binding pockets that enable them to bind in a highly specific way to complementary shapes on foreign molecules (shown in black). Binding to foreigh substances can neutralize them and also tag them to facilitate their removal by white blood cells.
As final example, consider the hormone insulin, which binds to specific pockets on insulin receptors embeded in the cell membranes of fat cells and muscle cells. Once again, binding causes a change in the conformation (shape) of the receptor protein which triggers a sequence of biochemical events that result in the insertion of glucose transporters (transmembrane transport proteins) into the plasma membrane as shown below.
As you will see in the next section, the structure of proteins enables them to serve a wide variety of functions.
Each of us has tens of thousands of proteins, which serve a variety of functions, and each protein has a unique three-dimensional structure that specifies its function. For example, hemoglobin is a protein found in red blood cells, which plays a key role in oxygen transport; it has 4 subunits of two distinct types (2 alpha and 2 beta subunits).
from http://gened.emc.maricopa.edu/bio/bio181/BIOBK/3_14d.jpg
|
Sickle Cell Anemia The critical relationship between protein structure and function is dramatically illustrated by sickle cell anemia, an inherited disease seen in people whose ancestors came from Africa, the Middle East, the Mediterranean, or India. In the U.S. about 4 out of every 1,000 African Americans has sickle cell disease (about 80,000 people), and about 10% carry the sickle cell trait. People with sickle cell disease an abnormal type of hemoglobin, called hemoglobin S (instead of normal hemoglobin A). Hemoglobin S differs from hemoglobin A in that the amino acid valine is found at position number 6 in the beta chain instead of the amino acid glutamate. Unlike glutamate, the side chain of valine is very non-polar and creates a sticky patch on the outside of each of the beta chains. There is a complementary sticky patch elsewhere on the hemoglobin, but it is masked as long as the hemoglobin molecules are bound to oxygen. However, if large numbers of hemoglobin molecules become deoxygenated, the sticky sites created by the abnormal valines begin to bind to the complementary sticky site on other hemoglobin molecules. This forms long aggregates of hemoglobin that distort the red blood cell and give it a characteristic sickle shape. This causes red cells to aggregate and impairs their ability to circulate through small blood vessels (arterioles and capillaries), and it also makes them fragile, shortening their life span and leading to anemia. |
Some proteins function as enzymes, i.e., proteins that catalyze specific biochemical reactions. Enzymes facilitate biochemical reactions and speed them up enormously, making them as much as a million times faster. There are thousands of enzymes, and each type facilitates a specific biochemical reaction. In other words, a given enzyme only acts on specific reactant molecules (substrates) to produce a specific end product or products. The diagram below illustrates enzymatic cleavage of the disaccharide lactose (the substrate) into the monosaccharides galactose and glucose.
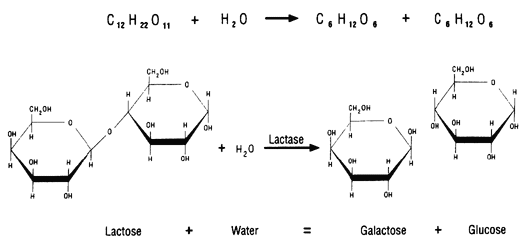
Source: http://www.indiana.edu/~ensiweb/lessons/tp.milk3.html
The three-dimensional shape of an enzyme will include a very specific binding site that the substrate will fit into very precisely, in much the same way that a key fits a specific lock.
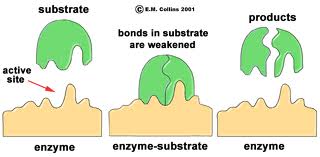
Once the substrate is bound the enzyme cleaves the substrate and the products are released. While this cartoon illustrates cleavage of a substrate, many enzymes synthesis new biochemicals by binding two substrates together to form a new product. A particular cell may have thousands of distinct enzymes catalyzing many different reactions.
The short video below illustrates the basics of how an enzyme works.
Sources: http://youtu.be/V4OPO6JQLOE
Biochemical reactions may require a whole series of steps, each of which is catalyzed by a separate enzyme. A good example is the series of reactions by which glucose is metabolized to create cellular energy in the form of ATP (adenosine triphosphate).
These reactions are illustrated in the figure below.
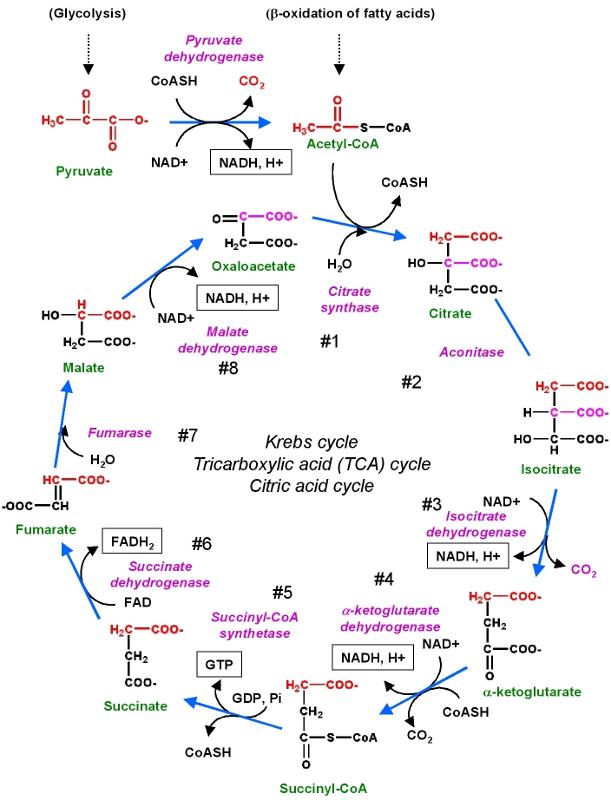
Source: http://chemwiki.ucdavis.edu/Core/Biological_Chemistry/Metabolism/Kreb's_Cycle
The illustration on the right shows the protein lysozyme (red, white, blue, and gray amino acids), which is an important defensive enzyme found in tears, saliva, and mucus. Lysozyme's function is to break down the polysaccharides (sugar polymers) that are components of bacterial cell walls. Initially, lysozyme is synthesized as a single long polypeptide chain, but it folds in a characteristic way to form a globular protein with a characteristic pocket. A bacterial polysaccharide (shown in green) binds to lysozyme because it fits precisely into the pocket in the same way that a key fits into a lock. Once this specific binding has occurred, lysozyme destroys the bacterial polysaccharide by cleaving it into pieces.
 Antibodies are defensive proteins that have binding sites whose three-dimensional structure allows them to identify and bind to very specific foreign molecules. By binding to foreign proteins they can help neutralize them and tag them, facilitating their engulfment and removal by defensive cells. IgG antibodies have a quaternary structure with four subunits, two "light chains" and two "heavy chains." The chains are bound to one another through disulfide bridges, shown to the right as "-S-S-" bonds. After birth, each B-lymphocyte can manufacture antibodies for only one specific foreign shape. The portion of an antigen that is specifically recognized by an antibody is referred to an an "epitope." In essence, the epitope is a particular portion of an antigen that has a distinctive molecular shape that fits into the protein binding site on an antibody.
Antibodies are defensive proteins that have binding sites whose three-dimensional structure allows them to identify and bind to very specific foreign molecules. By binding to foreign proteins they can help neutralize them and tag them, facilitating their engulfment and removal by defensive cells. IgG antibodies have a quaternary structure with four subunits, two "light chains" and two "heavy chains." The chains are bound to one another through disulfide bridges, shown to the right as "-S-S-" bonds. After birth, each B-lymphocyte can manufacture antibodies for only one specific foreign shape. The portion of an antigen that is specifically recognized by an antibody is referred to an an "epitope." In essence, the epitope is a particular portion of an antigen that has a distinctive molecular shape that fits into the protein binding site on an antibody.
Watch the short video below to see an illustration of antibody action. The beginning of the video shows red and white blood cells flowing through a blood vessel. The potato-shaped objects that you see next represent viruses that begin binding to receptors on a cell. The green Y-shaped objects represent antibodies that bind to the virus. Finally, the Medusa-like structure represents a white blood cell that engulfs the antibody-tagged virus and destroys it.
There are also structural proteins, which are frequently long and fibrous, such as silk, keratin in hair, and collagen in tendons and ligaments.
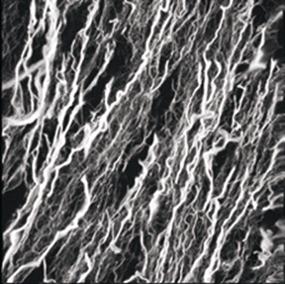
Source: http://www.sdsc.edu/ScienceAlive/reel6/collagen.gif
There are contractile proteins, such as actin and myosin, that provide movement in muscles and movement within single cells.
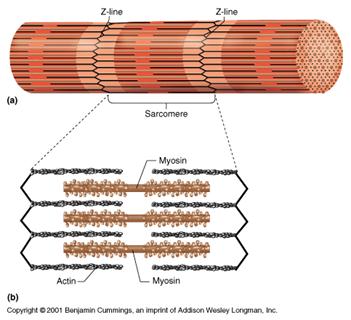
Source: http://www.bmb.psu.edu/courses/bisci004a/muscle/musc-img/myofibril.jpg
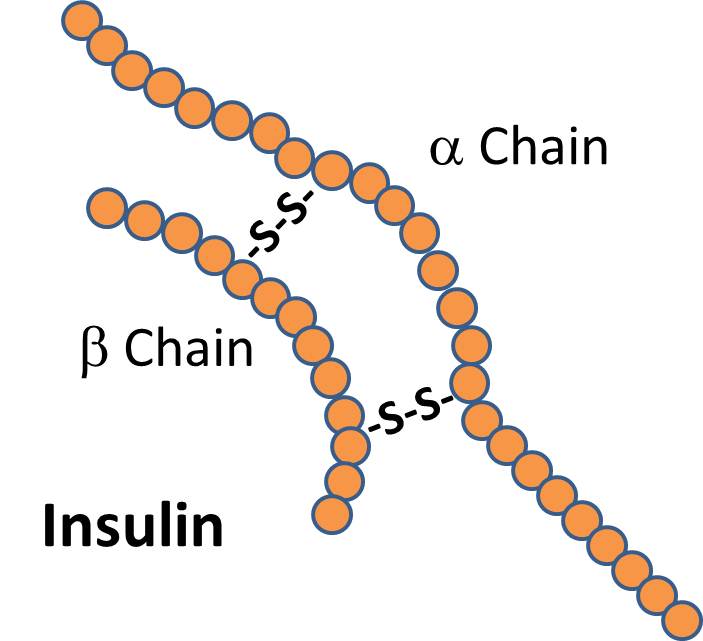 There are signal proteins, such as the hormone insulin, which consists of two polypeptide chains linked together with disulfide (two sulfur) bridges.
There are signal proteins, such as the hormone insulin, which consists of two polypeptide chains linked together with disulfide (two sulfur) bridges.
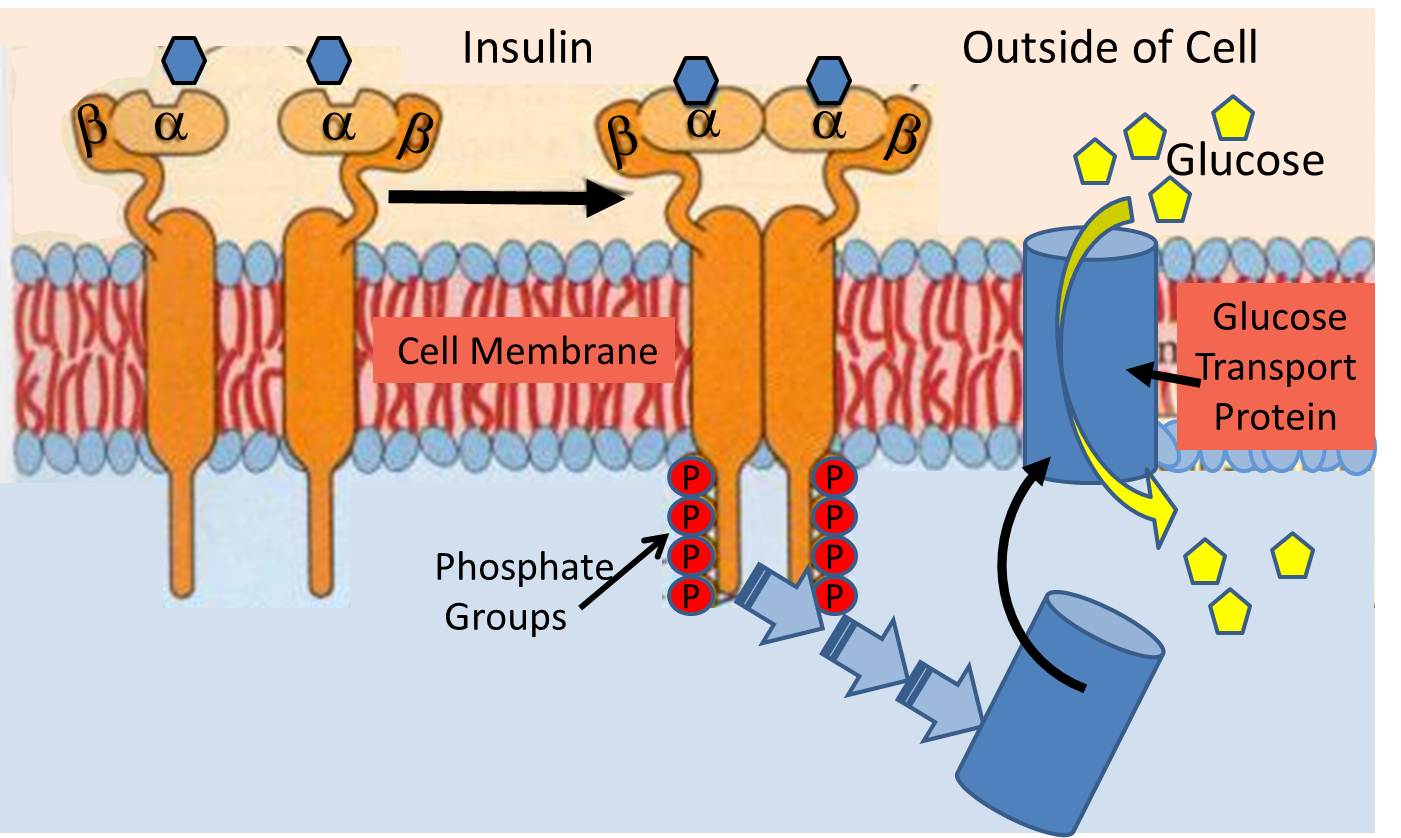 The insulin receptor (a recognition protein) is embedded in the cell membranes of muscle, fat cells and certain types of other cells. Its function is to facilitate their uptake of glucose from the blood stream through special glucose transport proteins that are normally present inside the cell in an inactive form. For example, in muscle cells, the glucose transporter is called "GLUT4". When the insulin molecule binds to the alpha subunits of the receptor, it triggers a chain reaction within the cytosol (the interior of the cell) that activates GLUT4 and causes it to be translocated and inserted into the cell membrane.
The insulin receptor (a recognition protein) is embedded in the cell membranes of muscle, fat cells and certain types of other cells. Its function is to facilitate their uptake of glucose from the blood stream through special glucose transport proteins that are normally present inside the cell in an inactive form. For example, in muscle cells, the glucose transporter is called "GLUT4". When the insulin molecule binds to the alpha subunits of the receptor, it triggers a chain reaction within the cytosol (the interior of the cell) that activates GLUT4 and causes it to be translocated and inserted into the cell membrane.
See http://arbl.cvmbs.colostate.edu/hbooks/pathphys/endocrine/pancreas/insulin_phys.html for a Flash model on insulin action.
With the exception of simple diffusion, proteins are also essential for moving polarized or charged molecules and large molecules across cell membranes.
Small molecules like oxygen and carbon dioxide can diffuse across the lipid bilayer of the cell membrane. The direction of movement depends on the concentration gradient. Substances with higher concentration inside the cell (e.g., CO2) will diffuse out of the cell toward the side with lower concentration. Substances in higher concentration outside the cell (e.g., O2) will diffuse to the inside of the cell, i.e., down the concentration gradient.
However, many other molecules cannot cross cell membranes by simple diffusion and require specialized mechanisms for movement across membranes. A variety of transport proteins, frequently aggregates of protein subunits, provide a way of transporting charged molecules and large molecules through one of two mechanisms:
Polar molecules and charged ions cannot cross the lipid bilayer; their transit relies on special transport channels created by proteins embedded in the cell membrane. Facilitated transport is passive in that it does not require expenditure of cellular energy, and as with simple diffusion, movement of the molecules is down a concentration gradient from high concentration to low concentration. There are specific proteins for each substance transported by this mechanism, and transit can be regulated by the cell. Molecules like glucose and amino acids are transported this way. They will bind to their carrier/transpport protein, and binding triggers a change in the shape of the carrier which moves the molecule across the membrane. Once the molecule is released, the carrier returns to its original shape (conformation).
Active transport also relies on transmembrane transport proteins, but this process is able to transport substances against a conentration gradient, meaning that even if the concentration of, say potassium ions, is higher inside the cell than outside, more potassium can be transported into the cell. This is because cellular energy (ATP) is expended.
Proteins, then, play an integral role in the function of a cell. Many are embedded in the cell's membranes or span the entire lipid bilayer where they play an important role in recognition, signaling, and transport.
An angstrom is one ten-millionth of a millimeter, or 1×10−10 meters. The illustration below gives an idea of the relative scale of some of the biological structures discussed above.
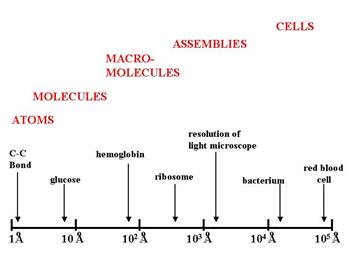
The distance between two carbon atoms in a fatty acid chain is a little over one angstrom. A glucose molecule is about 9 angstroms. Bacteria are tens of thousands of angstroms. And as a rough estimate, a typical human cell might be approximately 1/100th of a millimeter which is about 1/10th the width of a human hair. For an intriguing perspective on the size of things from the smallest to the largest objects in the universe, take a look at http://htwins.net/scale2/.
Despite their microscopic size, cells have a lot going on all the time. Diagrams and photomicrographs depict cells as rigid, static sacs that are frozen in time, but if we could somehow take a trip inside a cell, we would be staggered by the beauty, complexity, and incredible activity. You can get at least a glimpse of the inner life of a cell by watching the Harvard University animation, "The Inner Life of a Cell" (full length with narration), showing leukocyte activation in inflammation. Some of the terms used in the video will be foreign to you, but the video provides a magnificent sense of the inner works of cells, and it shows cells to be dynamic structures in which many processes are taking place continually.
One of the fundamental concepts in biology is that simple molecular structures (monomers) can be linked together to form increasingly complex structures. For example, monomers of sugars, such as glucose and fructose, can be linked together to form very large polysaccharides like starch and glycogen. Amino acids can be linked together to form polypeptides (proteins). Nucleotides can be linked together to form DNA and RNA.
In addition to linking molecules together to form long chains, many molecules will self-assemble under appropriate conditions to form increasingly complex molecular aggregates, such as membranes or lipoproteins. And biological membranes can provide the structure for intracellular organelles that can carry out specialized and complex functions. For example, microtubules are hollow cylinders that provide an internal scaffolding for eukaryotic cells and also provide tracks along which membrane-bound materials or organelles can be transported from place to place within the cell. For example, the microtubule network connects the Golgi apparatus with the plasma membrane to guide secretory vesicles for export or for insertion into the plasma membrane. The movement of these membrane-bound 'packages' along microtubules is facilitated by motor proteins (the carriers) which move along the microtubule by changing their three-dimensional conformation. This process is powered by adenosine triphosphate (ATP). With each 'step', the motor molecule releases one portion of the microtubule and grips a second site farther long the filament.
These microtubules are polymers composed of subunits of a protein called tubulin. Each subunit of the microtubule is made of two slightly different but closely related simpler units called alpha-tubulin (shown in the figure below as yellow beads) and beta-tubulin (shown as green beads). Under appropriate conditions these subunits aggregate or self-assemble in a particular way that rapidly forms a microtubule. Conversely, these microtubules can also rapidly disaggregate.
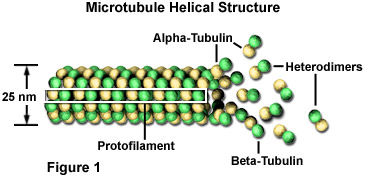
Source: https://micro.magnet.fsu.edu/cells/microtubules/microtubules.html
The video below is a TED talk in which animator David Bolinsky describes a collaboration between animators and biologists at Harvard University in which one gains a vision of the beauty and complexity of eukaryotic cells. Note that the phenomenon of monomers being assembled into complex and highly functional macromolecular polymers is illustrated in several places.
The entire talk is 9:49. Advance the video to 3:24 to skip the introductory description. The real action begins at about 6:50. There is nothing you have to memorize in this. Just appreciate the complexity and beauty of cells.
The next video below provides a basic overview of cell structure and function (6:00 min.), and the second provides a short description of the structure and function of the organelles in a eukaryotic cell (4:46 min.).
One can see that cells, the smallest unit that meets the criteria for being alive, are highly complex. Nevertheless, this complexity results from simple molecules being linked together to form a myriad of increasingly diverse and complex structures, and these, in turn, provide the basis for an even higher level of organization and complexity by assembling into macromolecular complexes, such as membranes, organelles, microtubules, and lipoproteins. From the cellular level, one can then envision the aggregation of cells to form tissues, which become the basis for organs and even organ systems in an incredibly diverse array of multicellular organisms.
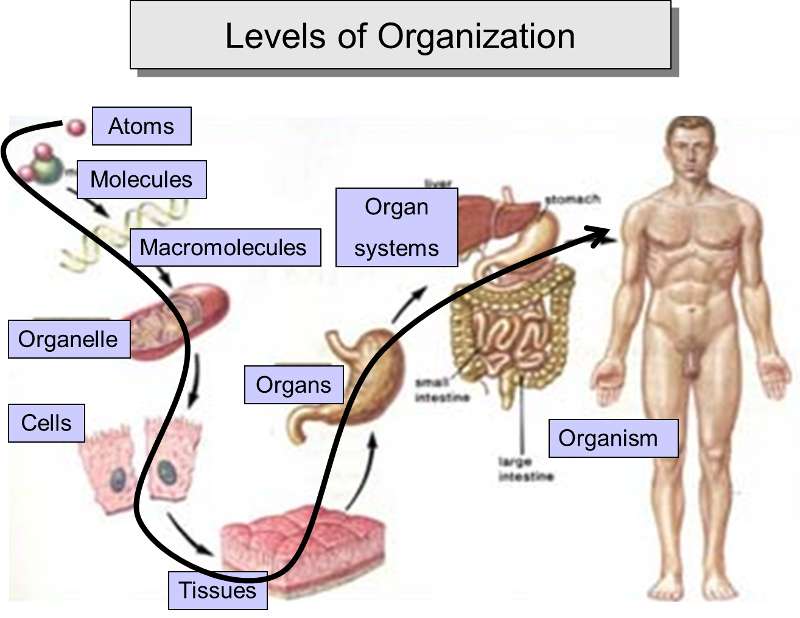
Adapted from: http://www.theorganicstartupbook.com/2012/07/07/evolutionary-levels-sublevels-4-of-5/